[English] 日本語
 Yorodumi
Yorodumi- PDB-1qjs: mammalian blood serum haemopexin glycosylated-native protein and ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1qjs | ||||||
|---|---|---|---|---|---|---|---|
| Title | mammalian blood serum haemopexin glycosylated-native protein and in complex with its ligand haem | ||||||
 Components Components | HEMOPEXIN | ||||||
 Keywords Keywords |  TRANSPORT PROTEIN / HAEM BINDING PROTEIN / TRANSPORT PROTEIN / HAEM BINDING PROTEIN /  BETA PROPELLER / HAEM BINDING AND TRANSPORT / BETA PROPELLER / HAEM BINDING AND TRANSPORT /  IRON METABOLISM IRON METABOLISM | ||||||
| Function / homology |  Function and homology information Function and homology informationheme transmembrane transporter activity / intracellular iron ion homeostasis /  heme binding / extracellular region / heme binding / extracellular region /  metal ion binding metal ion bindingSimilarity search - Function | ||||||
| Biological species |   ORYCTOLAGUS CUNICULUS (rabbit) ORYCTOLAGUS CUNICULUS (rabbit) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SIR / Resolution: 2.9 Å SIR / Resolution: 2.9 Å | ||||||
 Authors Authors | Paoli, M. / Baker, H.M. / Morgan, W.T. / Smith, A. / Baker, E.N. | ||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 1999 Journal: Nat.Struct.Biol. / Year: 1999Title: Crystal Structure of Hemopexin Reveals a Novel High-Affinity Heme Site Formed between Two Beta-Propeller Domains. Authors: Paoli, M. / Anderson, B.F. / Baker, H.M. / Morgan, W.T. / Smith, A. / Baker, E.N. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1qjs.cif.gz 1qjs.cif.gz | 180.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1qjs.ent.gz pdb1qjs.ent.gz | 141.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1qjs.json.gz 1qjs.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qj/1qjs https://data.pdbj.org/pub/pdb/validation_reports/qj/1qjs ftp://data.pdbj.org/pub/pdb/validation_reports/qj/1qjs ftp://data.pdbj.org/pub/pdb/validation_reports/qj/1qjs | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1qhuC  1fblS 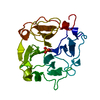 1hxnS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (0.43434, -0.90066, 0.01304), Vector  : : |
- Components
Components
| #1: Protein |  Mass: 51832.293 Da / Num. of mol.: 2 / Fragment: BETA-PROPELLER DOMAIN, HAEM LIGAND / Source method: isolated from a natural source Details: COVALENT LINK BETWEEN FE OF HAEM LIGAND AND (I) NE2 OF HIS 213 AND (II) NE2 OF HIS 266 Source: (natural)   ORYCTOLAGUS CUNICULUS (rabbit) / Tissue: BLOOD SERUM / References: UniProt: P20058 ORYCTOLAGUS CUNICULUS (rabbit) / Tissue: BLOOD SERUM / References: UniProt: P20058#2: Chemical |  Heme B Heme B#3: Chemical |  Phosphate Phosphate#4: Chemical | ChemComp-CL /  Chloride Chloride#5: Chemical | ChemComp-NA / Compound details | THIS ENTRY ACCOMPANIES PDB ENTRY 1QHU, WHICH IS FOR DEGLYCOSYLATED HAEMOPEXIN. THE CRYSTALS WERE ...THIS ENTRY ACCOMPANIE | Sequence details | THE N-TERMINUS (-25 TO 24 IN PDB NUMBERING, 1 TO 48 IN SWS) IS DISORDERED IN THE ELECTRON DENSITY ...THE N-TERMINUS (-25 TO 24 IN PDB NUMBERING, 1 TO 48 IN SWS) IS DISORDERED | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 2 X-RAY DIFFRACTION / Number of used crystals: 2 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.04 Å3/Da / Density % sol: 40 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop / pH: 7.9 Details: HANGING DROP, 4 DEGREES CENTIGRADE. RESEVOIR SOLUTION: 15-22% PEG 6000, 0.1-0.2 M TRIS HCL PH 7.9, 0.05-0.1 M EDTA, 0.05-0.2 NACL, PROTEIN COMPLEX SOLUTION: 65 MG/ML IN 0.01 M TRIS PH 7.9, 0. ...Details: HANGING DROP, 4 DEGREES CENTIGRADE. RESEVOIR SOLUTION: 15-22% PEG 6000, 0.1-0.2 M TRIS HCL PH 7.9, 0.05-0.1 M EDTA, 0.05-0.2 NACL, PROTEIN COMPLEX SOLUTION: 65 MG/ML IN 0.01 M TRIS PH 7.9, 0.01 M NACL, 0.1 M EDTA | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 4 ℃ / Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Wavelength: 1.5418 ROTATING ANODE / Wavelength: 1.5418 |
| Detector | Type: RIGAKU IMAGE PLATE / Detector: IMAGE PLATE / Date: Jun 15, 1997 / Details: COLLIMATOR |
| Radiation | Monochromator: GRAPHITE(002) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1.5418 Å / Relative weight: 1 : 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.9→20 Å / Num. obs: 20915 / % possible obs: 97.5 % / Redundancy: 2.2 % / Biso Wilson estimate: 44 Å2 / Rmerge(I) obs: 0.064 / Net I/σ(I): 6.2 |
| Reflection shell | Resolution: 2.9→3.05 Å / Redundancy: 2 % / Rmerge(I) obs: 0.251 / Mean I/σ(I) obs: 2.2 / % possible all: 87.5 |
| Reflection shell | *PLUS % possible obs: 87.5 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  SIR SIRStarting model: 1HXN AND 1FBL Resolution: 2.9→20 Å / Cross valid method: THROUGHOUT / σ(F): 2.5 Details: ELECTRON DENSITY FOR RESIDUES ARG 214, SER 215, HIS 223 AND C-TERMINAL HIS 435 IS POORLY DEFINED. FOR BOTH CHAINS A AND B RESIDUES 219 - 222 COULD NOT BE FITTED INTO THE ELECTRON DENSITY ...Details: ELECTRON DENSITY FOR RESIDUES ARG 214, SER 215, HIS 223 AND C-TERMINAL HIS 435 IS POORLY DEFINED. FOR BOTH CHAINS A AND B RESIDUES 219 - 222 COULD NOT BE FITTED INTO THE ELECTRON DENSITY MAPS AND ARE NOT INCLUDED IN THE MODEL.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 39 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.9→20 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj








