[English] 日本語
 Yorodumi
Yorodumi- PDB-1qhu: MAMMALIAN BLOOD SERUM HAEMOPEXIN DEGLYCOSYLATED AND IN COMPLEX WI... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1qhu | ||||||
|---|---|---|---|---|---|---|---|
| Title | MAMMALIAN BLOOD SERUM HAEMOPEXIN DEGLYCOSYLATED AND IN COMPLEX WITH ITS LIGAND HAEM | ||||||
 Components Components | PROTEIN (HEMOPEXIN) | ||||||
 Keywords Keywords |  BINDING PROTEIN / BINDING PROTEIN /  BETA PROPELLER / HAEM BINDING AND TRANSPORT / BETA PROPELLER / HAEM BINDING AND TRANSPORT /  IRON METABOLISM IRON METABOLISM | ||||||
| Function / homology |  Function and homology information Function and homology informationheme transmembrane transporter activity / intracellular iron ion homeostasis /  heme binding / extracellular region / heme binding / extracellular region /  metal ion binding metal ion bindingSimilarity search - Function | ||||||
| Biological species |   Oryctolagus cuniculus (rabbit) Oryctolagus cuniculus (rabbit) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.3 Å MOLECULAR REPLACEMENT / Resolution: 2.3 Å | ||||||
 Authors Authors | Paoli, M. / Baker, H.M. / Morgan, W.T. / Smith, A. / Baker, E.N. | ||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 1999 Journal: Nat.Struct.Biol. / Year: 1999Title: Crystal structure of hemopexin reveals a novel high-affinity heme site formed between two beta-propeller domains. Authors: Paoli, M. / Anderson, B.F. / Baker, H.M. / Morgan, W.T. / Smith, A. / Baker, E.N. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1qhu.cif.gz 1qhu.cif.gz | 105.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1qhu.ent.gz pdb1qhu.ent.gz | 76.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1qhu.json.gz 1qhu.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qh/1qhu https://data.pdbj.org/pub/pdb/validation_reports/qh/1qhu ftp://data.pdbj.org/pub/pdb/validation_reports/qh/1qhu ftp://data.pdbj.org/pub/pdb/validation_reports/qh/1qhu | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1qjsC  1fblS 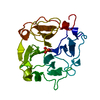 1hxnS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
-Protein , 1 types, 1 molecules A
| #1: Protein | Mass: 51832.293 Da / Num. of mol.: 1 / Fragment: BETA-PROPELLER DOMAIN / Source method: isolated from a natural source Details: COVALENT LINK BETWEEN FE OF HAEM LIGAND AND I) NE2 OF HIS 213 AND II) NE2 OF HIS 265 Source: (natural)   Oryctolagus cuniculus (rabbit) / Tissue: BLOOD SERUM / References: UniProt: P20058 Oryctolagus cuniculus (rabbit) / Tissue: BLOOD SERUM / References: UniProt: P20058 |
|---|
-Non-polymers , 5 types, 218 molecules 








| #2: Chemical |  Chloride Chloride#3: Chemical | ChemComp-NA / #4: Chemical | ChemComp-PO4 / |  Phosphate Phosphate#5: Chemical | ChemComp-HEM / |  Heme B Heme B#6: Water | ChemComp-HOH / |  Water Water |
|---|
-Details
| Compound details | PROTEIN WAS DEGLYSOSYL| Nonpolymer details | LIGAND COMPLEXED TO HAEMOPEXIN ION BOUND IN THE CENTRAL TUNNEL OF THE C-TERMINAL DOMAIN IONS BOUND ...LIGAND COMPLEXED TO HAEMOPEXIN | Sequence details | N-TERMINUS IS DISORDERED IN ELECTRON DENSITY MAPS NUMERING OF RESIDUES IS SUCH THAT IT IS ...N-TERMINUS IS DISORDERED | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 2 X-RAY DIFFRACTION / Number of used crystals: 2 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.92 Å3/Da / Density % sol: 35 % | ||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | pH: 7.5 Details: HANGING DROP, 4 DEGREES CENTIGRADES, RESEVOIR SOLUTION: 19-22% PEG 4000, 0.05 M TRIS HCL PH 7.5, 0.05-0.5 M EDTA, 0.2 NACL PROTEIN COMPLEX SOLUTION: 40 MG/ ML IN 0.05 M TRIS PH 7.0 AND 0.2 M NACL | ||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 4 ℃ / pH: 7 / Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRL SSRL  / Beamline: BL7-1 / Wavelength: 1.08 / Beamline: BL7-1 / Wavelength: 1.08 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Feb 1, 1997 / Details: COLLIMATOR |
| Radiation | Monochromator: CYCLINDRICALLY BENT TRIANGULAR SI(111) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1.08 Å / Relative weight: 1 : 1.08 Å / Relative weight: 1 |
| Reflection | Resolution: 2.3→20 Å / Num. obs: 19233 / % possible obs: 94.7 % / Redundancy: 2.4 % / Biso Wilson estimate: 38 Å2 / Rmerge(I) obs: 0.09 / Net I/σ(I): 6 |
| Reflection shell | Resolution: 2.3→2.42 Å / Redundancy: 2.1 % / Rmerge(I) obs: 0.324 / Mean I/σ(I) obs: 2.1 / % possible all: 77 |
| Reflection | *PLUS Rmerge(I) obs: 0.09 |
| Reflection shell | *PLUS % possible obs: 77 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1HXN AND 1FBL Resolution: 2.3→20 Å / Cross valid method: THROUGHOUT / σ(F): 2.5 / ESU R: 1.29 / ESU R Free: 0.33 Details: ELECTRON DENSITY FOR RESIDUES ARG 214, SER 215 AND HIS 222 IS POORLY DEFINED
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 35 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.3→20 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: REFMAC / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 2.3 Å / Lowest resolution: 20 Å / σ(F): 2.5 / % reflection Rfree: 5 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS Biso mean: 35 Å2 |
 Movie
Movie Controller
Controller












 PDBj
PDBj




