[English] 日本語
 Yorodumi
Yorodumi- PDB-1mea: METHIONYL-TRNA SYNTHETASE ZINC BINDING DOMAIN. 3D STRUCTURE AND H... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1mea | ||||||
|---|---|---|---|---|---|---|---|
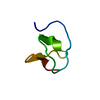
| Title | METHIONYL-TRNA SYNTHETASE ZINC BINDING DOMAIN. 3D STRUCTURE AND HOMOLOGY WITH RUBREDOXIN AND GAG RETROVIRAL PROTEINS | ||||||
 Components Components | METHIONYL-tRNA SYNTHETASE Methionine—tRNA ligase Methionine—tRNA ligase | ||||||
 Keywords Keywords | AMINOACYL-TRNA SYNTHASE | ||||||
| Function / homology |  Function and homology information Function and homology information methionine-tRNA ligase / methionine-tRNA ligase /  methionine-tRNA ligase activity / methionyl-tRNA aminoacylation / aminoacyl-tRNA synthetase multienzyme complex / methionine-tRNA ligase activity / methionyl-tRNA aminoacylation / aminoacyl-tRNA synthetase multienzyme complex /  tRNA binding / protein homodimerization activity / zinc ion binding / tRNA binding / protein homodimerization activity / zinc ion binding /  ATP binding / ATP binding /  membrane / membrane /  cytosol cytosolSimilarity search - Function | ||||||
| Biological species |   Escherichia coli (E. coli) Escherichia coli (E. coli) | ||||||
| Method |  SOLUTION NMR SOLUTION NMR | ||||||
 Authors Authors | Fourmy, D. / Dardel, F. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1993 Journal: J.Mol.Biol. / Year: 1993Title: Methionyl-tRNA synthetase zinc binding domain. Three-dimensional structure and homology with rubredoxin and gag retroviral proteins. Authors: Fourmy, D. / Dardel, F. / Blanquet, S. #1:  Journal: J.Mol.Biol. / Year: 1993 Journal: J.Mol.Biol. / Year: 1993Title: Mapping of the Zinc Binding Domain of Escherichia Coli Methionyl-tRNA Synthetase Authors: Fourmy, D. / Meinnel, T. / Mechulam, Y. / Blanquet, S. #2:  Journal: J.Mol.Biol. / Year: 1990 Journal: J.Mol.Biol. / Year: 1990Title: Crystallographic Study at 2.5 Angstroms Resolution of the Interaction of Methionyl-tRNA Synthetase from Escherichia Coli with ATP Authors: Brunie, S. / Zelwer, C. / Risler, J.-L. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1mea.cif.gz 1mea.cif.gz | 17 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1mea.ent.gz pdb1mea.ent.gz | 9.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1mea.json.gz 1mea.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/me/1mea https://data.pdbj.org/pub/pdb/validation_reports/me/1mea ftp://data.pdbj.org/pub/pdb/validation_reports/me/1mea ftp://data.pdbj.org/pub/pdb/validation_reports/me/1mea | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| Atom site foot note | 1: CYS 10 - PRO 11 OMEGA ANGLE = 216.960 PEPTIDE BOND DEVIATES SIGNIFICANTLY FROM TRANS CONFORMATION 2: SER 15 - PRO 16 OMEGA ANGLE = 216.723 PEPTIDE BOND DEVIATES SIGNIFICANTLY FROM TRANS CONFORMATION | |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide |  Methionine—tRNA ligase Methionine—tRNA ligaseMass: 2969.311 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Escherichia coli (E. coli) / Gene: GAG / Gene (production host): GAG / References: UniProt: P00959, Escherichia coli (E. coli) / Gene: GAG / Gene (production host): GAG / References: UniProt: P00959,  methionine-tRNA ligase methionine-tRNA ligase |
|---|---|
| #2: Chemical | ChemComp-ZN / |
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR |
|---|
- Processing
Processing
| Software |
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| NMR ensemble | Conformers submitted total number: 1 |
 Movie
Movie Controller
Controller










 PDBj
PDBj


