+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ls6 | ||||||
|---|---|---|---|---|---|---|---|
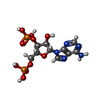
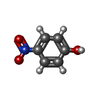
| Title | Human SULT1A1 complexed with PAP and p-Nitrophenol | ||||||
 Components Components | aryl sulfotransferase | ||||||
 Keywords Keywords |  TRANSFERASE / sult 1A1 / PAP / TRANSFERASE / sult 1A1 / PAP /  p-Nitrophenol / positive cooperativity / two substrate binding sites p-Nitrophenol / positive cooperativity / two substrate binding sites | ||||||
| Function / homology |  Function and homology information Function and homology information flavonol 3-sulfotransferase activity / flavonol 3-sulfotransferase activity /  aryl sulfotransferase / aryl sulfotransferase /  steroid sulfotransferase activity / 3'-phosphoadenosine 5'-phosphosulfate binding / steroid sulfotransferase activity / 3'-phosphoadenosine 5'-phosphosulfate binding /  aryl sulfotransferase activity / Cytosolic sulfonation of small molecules / flavonoid metabolic process / 3'-phosphoadenosine 5'-phosphosulfate metabolic process / aryl sulfotransferase activity / Cytosolic sulfonation of small molecules / flavonoid metabolic process / 3'-phosphoadenosine 5'-phosphosulfate metabolic process /  sulfation / ethanol catabolic process ... sulfation / ethanol catabolic process ... flavonol 3-sulfotransferase activity / flavonol 3-sulfotransferase activity /  aryl sulfotransferase / aryl sulfotransferase /  steroid sulfotransferase activity / 3'-phosphoadenosine 5'-phosphosulfate binding / steroid sulfotransferase activity / 3'-phosphoadenosine 5'-phosphosulfate binding /  aryl sulfotransferase activity / Cytosolic sulfonation of small molecules / flavonoid metabolic process / 3'-phosphoadenosine 5'-phosphosulfate metabolic process / aryl sulfotransferase activity / Cytosolic sulfonation of small molecules / flavonoid metabolic process / 3'-phosphoadenosine 5'-phosphosulfate metabolic process /  sulfation / ethanol catabolic process / sulfation / ethanol catabolic process /  sulfotransferase activity / catecholamine metabolic process / Paracetamol ADME / amine metabolic process / estrogen metabolic process / xenobiotic metabolic process / sulfotransferase activity / catecholamine metabolic process / Paracetamol ADME / amine metabolic process / estrogen metabolic process / xenobiotic metabolic process /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 1.9 Å MOLECULAR REPLACEMENT / Resolution: 1.9 Å | ||||||
 Authors Authors | Gamage, N.U. / Barnett, A.C. / Tresillian, M. / Latham, C.F. / Liyou, N.E. / McManus, M.E. / Martin, J.L. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2003 Journal: J.Biol.Chem. / Year: 2003Title: Structure of a human carcinogen-converting enzyme, SULT1A1. Structural and kinetic implications of substrate inhibition. Authors: Gamage, N.U. / Duggleby, R.G. / Barnett, A.C. / Tresillian, M. / Latham, C.F. / Liyou, N.E. / McManus, M.E. / Martin, J.L. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ls6.cif.gz 1ls6.cif.gz | 76.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ls6.ent.gz pdb1ls6.ent.gz | 57.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ls6.json.gz 1ls6.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ls/1ls6 https://data.pdbj.org/pub/pdb/validation_reports/ls/1ls6 ftp://data.pdbj.org/pub/pdb/validation_reports/ls/1ls6 ftp://data.pdbj.org/pub/pdb/validation_reports/ls/1ls6 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1aquS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein |  Mass: 34220.309 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: SULT1A1 / Organ: liver Homo sapiens (human) / Gene: SULT1A1 / Organ: liver / Plasmid: PET20B / Species (production host): Escherichia coli / Production host: / Plasmid: PET20B / Species (production host): Escherichia coli / Production host:   Escherichia coli BL21(DE3) (bacteria) / Strain (production host): BL21(DE3) / References: UniProt: P50225, Escherichia coli BL21(DE3) (bacteria) / Strain (production host): BL21(DE3) / References: UniProt: P50225,  aryl sulfotransferase aryl sulfotransferase | ||
|---|---|---|---|
| #2: Chemical | ChemComp-A3P /  Adenosine 3',5'-bisphosphate Adenosine 3',5'-bisphosphate | ||
| #3: Chemical |  4-Nitrophenol 4-Nitrophenol#4: Water | ChemComp-HOH / |  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.79 Å3/Da / Density % sol: 56 % | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 8 Details: 20% PEG 4000, 0.1M Tris, 10 mM DTT, pH 8.0, VAPOR DIFFUSION, HANGING DROP, temperature 293K | ||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 20 ℃ / Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 290 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RUH3R / Wavelength: 1.5418 Å ROTATING ANODE / Type: RIGAKU RUH3R / Wavelength: 1.5418 Å |
| Detector | Type: RIGAKU RAXIS IV++ / Detector: IMAGE PLATE / Date: Jul 31, 2001 / Details: Osmic Maxflux confocal Blue |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1.5418 Å / Relative weight: 1 : 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 1.9→19.89 Å / Num. all: 32548 / Num. obs: 31483 / % possible obs: 96.7 % / Observed criterion σ(I): 0 / Redundancy: 2.6 % / Biso Wilson estimate: 18.5 Å2 / Rmerge(I) obs: 0.086 / Rsym value: 0.041 / Net I/σ(I): 8.6 |
| Reflection shell | Resolution: 1.9→2.02 Å / Redundancy: 1.35 % / Rmerge(I) obs: 0.245 / Mean I/σ(I) obs: 2.4 / Num. unique all: 4131 / Rsym value: 0.22 / % possible all: 86 |
| Reflection | *PLUS % possible obs: 91.4 % / Num. measured all: 82261 / Rmerge(I) obs: 0.041 |
| Reflection shell | *PLUS % possible obs: 63.8 % / Rmerge(I) obs: 0.22 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ID 1AQU Resolution: 1.9→19.88 Å / Rfactor Rfree error: 0.004 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: Flat model / Bsol: 62.3 Å2 / ksol: 0.35 e/Å3 | ||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 24.9 Å2
| ||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.9→19.88 Å
| ||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.9→2.02 Å / Rfactor Rfree error: 0.012 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 1.9 Å / Lowest resolution: 20 Å / % reflection Rfree: 10 % / Rfactor Rfree : 0.205 / Rfactor Rwork : 0.205 / Rfactor Rwork : 0.182 : 0.182 | ||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller













 PDBj
PDBj