+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ega | ||||||
|---|---|---|---|---|---|---|---|
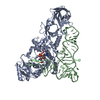
| Title | CRYSTAL STRUCTURE OF A WIDELY CONSERVED GTPASE ERA | ||||||
 Components Components | PROTEIN (GTP-BINDING PROTEIN ERA) | ||||||
 Keywords Keywords |  HYDROLASE / ERA / HYDROLASE / ERA /  GTPASE / RNA-BINDING / RAS-LIKE GTPASE / RNA-BINDING / RAS-LIKE | ||||||
| Function / homology |  Function and homology information Function and homology information guanosine tetraphosphate binding / guanosine tetraphosphate binding /  ribosomal small subunit binding / ribosomal small subunit binding /  ribosomal small subunit biogenesis / small ribosomal subunit rRNA binding / ribosomal small subunit biogenesis / small ribosomal subunit rRNA binding /  ribosomal small subunit assembly / ribosomal small subunit assembly /  rRNA binding / rRNA binding /  protein phosphorylation / protein phosphorylation /  GTPase activity / GTP binding / GTPase activity / GTP binding /  RNA binding ... RNA binding ... guanosine tetraphosphate binding / guanosine tetraphosphate binding /  ribosomal small subunit binding / ribosomal small subunit binding /  ribosomal small subunit biogenesis / small ribosomal subunit rRNA binding / ribosomal small subunit biogenesis / small ribosomal subunit rRNA binding /  ribosomal small subunit assembly / ribosomal small subunit assembly /  rRNA binding / rRNA binding /  protein phosphorylation / protein phosphorylation /  GTPase activity / GTP binding / GTPase activity / GTP binding /  RNA binding / RNA binding /  plasma membrane / plasma membrane /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Escherichia coli (E. coli) Escherichia coli (E. coli) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MIR / Resolution: 2.4 Å MIR / Resolution: 2.4 Å | ||||||
 Authors Authors | Chen, X. / Ji, X. | ||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 1999 Journal: Proc Natl Acad Sci U S A / Year: 1999Title: Crystal structure of ERA: a GTPase-dependent cell cycle regulator containing an RNA binding motif. Authors: X Chen / D L Court / X Ji /  Abstract: ERA forms a unique family of GTPase. It is widely conserved and essential in bacteria. ERA functions in cell cycle control by coupling cell division with growth rate. ERA homologues also are found in ...ERA forms a unique family of GTPase. It is widely conserved and essential in bacteria. ERA functions in cell cycle control by coupling cell division with growth rate. ERA homologues also are found in eukaryotes. Here we report the crystal structure of ERA from Escherichia coli. The structure has been determined at 2.4-A resolution. It reveals a two-domain arrangement of the molecule: an N-terminal domain that resembles p21 Ras and a C-terminal domain that is unique. Structure-based topological search of the C domain fails to reveal any meaningful match, although sequence analysis suggests that it contains a KH domain. KH domains are RNA binding motifs that usually occur in tandem repeats and exhibit low sequence similarity except for the well-conserved segment VIGxxGxxIK. We have identified a betaalphaalphabeta fold that contains the VIGxxGxxIK sequence and is shared by the C domain of ERA and the KH domain. We propose that this betaalphaalphabeta fold is the RNA binding motif, the minimum structural requirement for RNA binding. ERA dimerizes in crystal. The dimer formation involves a significantly distorted switch II region, which may shed light on how ERA protein regulates downstream events. #1:  Journal: Mol.Microbiol. / Year: 1998 Journal: Mol.Microbiol. / Year: 1998Title: Cell Cycle Arrest in Era GTPase Mutants: A Potential Growth Rate-Regulated Checkpoint in E. Coli Authors: Britton, R.A. / Powell, B.S. / Dasgupta, S. / Sun, Q. / Margolin, W. / Lupski, J.R. / Court, D.L. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ega.cif.gz 1ega.cif.gz | 128.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ega.ent.gz pdb1ega.ent.gz | 101 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ega.json.gz 1ega.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/eg/1ega https://data.pdbj.org/pub/pdb/validation_reports/eg/1ega ftp://data.pdbj.org/pub/pdb/validation_reports/eg/1ega ftp://data.pdbj.org/pub/pdb/validation_reports/eg/1ega | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (-0.436003, 0.021836, 0.89968), Vector  : : |
- Components
Components
| #1: Protein | Mass: 33858.078 Da / Num. of mol.: 2 / Source method: isolated from a natural source / Source: (natural)   Escherichia coli (E. coli) / Cellular location: CYTOPLASM/MEMBRANE / Strain: TAP106 / References: UniProt: P06616 Escherichia coli (E. coli) / Cellular location: CYTOPLASM/MEMBRANE / Strain: TAP106 / References: UniProt: P06616#2: Chemical | ChemComp-SO4 /  Sulfate Sulfate#3: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.5 Å3/Da / Density % sol: 65 % Description: THE STRUCTURE WAS SOLVED BY A COMBINATION OF MIR AND MAD METHODS. MAD DATA SETS, USING HG ANOMALOUS SCATTERING AT WAVELENGTH 1.00870A, 1.00764A, 0.99184A AND 1.00903A, WERE COLLECTED AT ...Description: THE STRUCTURE WAS SOLVED BY A COMBINATION OF MIR AND MAD METHODS. MAD DATA SETS, USING HG ANOMALOUS SCATTERING AT WAVELENGTH 1.00870A, 1.00764A, 0.99184A AND 1.00903A, WERE COLLECTED AT NSLS BEAMLINE X9B WITH MAR345 DETECTOR. | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | pH: 8 Details: PROTEIN 8MG/ML TRIS BUFFER 0.1M, PH 8.0 LITHIUM SULFATE 0.8M SODIUM CHLORIDE 0.8M | ||||||||||||||||||||||||||||||
| Components of the solutions |
| ||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Jun 18, 1998 / Details: BENT MIRRORS |
| Radiation | Monochromator: GRAPHITE / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1.5418 Å / Relative weight: 1 : 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.4→20 Å / Num. obs: 35696 / % possible obs: 99.7 % / Observed criterion σ(I): 0 / Redundancy: 6.4 % / Biso Wilson estimate: 53.4 Å2 / Rsym value: 0.056 / Net I/σ(I): 31.8 |
| Reflection shell | Resolution: 2.4→2.5 Å / Redundancy: 6.1 % / Mean I/σ(I) obs: 3.9 / Rsym value: 0.49 / % possible all: 99.1 |
| Reflection | *PLUS Rmerge(I) obs: 0.056 |
| Reflection shell | *PLUS % possible obs: 99.1 % / Rmerge(I) obs: 0.49 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MIR / Resolution: 2.4→8 Å / Rfactor Rfree error: 0.0086 / Data cutoff high absF: 100000 / Data cutoff low absF: 0.1 / Cross valid method: THROUGHOUT / σ(F): 4 MIR / Resolution: 2.4→8 Å / Rfactor Rfree error: 0.0086 / Data cutoff high absF: 100000 / Data cutoff low absF: 0.1 / Cross valid method: THROUGHOUT / σ(F): 4 Details: N-TERMINAL RESIDUES 1-3 FOR BOTH CHAINS, AND C-TERMINAL RESIDUES 296-301 FOR CHAIN A AND RESIDUES 297-301 FOR CHAIN B WERE NOT OBSERVED IN THE ELECTRON DNESITY AND NOT INCLUDED IN THE MODEL; ...Details: N-TERMINAL RESIDUES 1-3 FOR BOTH CHAINS, AND C-TERMINAL RESIDUES 296-301 FOR CHAIN A AND RESIDUES 297-301 FOR CHAIN B WERE NOT OBSERVED IN THE ELECTRON DNESITY AND NOT INCLUDED IN THE MODEL; RESIDUES 224 - 228 IN BOTH CHAINS WERE NOT OBSERVED IN THE ELECTRON DENSITY AND NOT INCLUDED IN THE REFINEMENT, BUT BUILT STEREOCHEMICALLY AND INCLUDED IN THE MODEL; RESTRAINTS BETWEEN THE TWO COPIES OF MOLECULES WERE APPLIED DURING THE INITIAL STAGE OF THE REFINEMENT ONLY
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 49.5 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.4→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.4→2.5 Å / Rfactor Rfree error: 0.035 / Total num. of bins used: 8
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.85 / Classification: refinement X-PLOR / Version: 3.85 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller










 PDBj
PDBj


