[English] 日本語
 Yorodumi
Yorodumi- PDB-1eaz: Crystal structure of the phosphoinositol (3,4)-bisphosphate bindi... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1eaz | ||||||
|---|---|---|---|---|---|---|---|
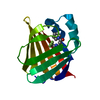
| Title | Crystal structure of the phosphoinositol (3,4)-bisphosphate binding PH domain of TAPP1 from human. | ||||||
 Components Components | TANDEM PH DOMAIN CONTAINING PROTEIN-1 | ||||||
 Keywords Keywords | LIPID BINDING PROTEIN / LIPID-BINDING PROTEIN / LIPID DEGRADATION /  PH DOMAIN / PHOSPHATIDYLINOSITOL (3 / 4)-BISPHOSPHATE / PH DOMAIN / PHOSPHATIDYLINOSITOL (3 / 4)-BISPHOSPHATE /  SIGNALLING SIGNALLING | ||||||
| Function / homology |  Function and homology information Function and homology information : / : /  luteinization / Leydig cell differentiation / ruffle organization / phosphatidylinositol biosynthetic process / phosphatidylinositol-3,4-bisphosphate binding / luteinization / Leydig cell differentiation / ruffle organization / phosphatidylinositol biosynthetic process / phosphatidylinositol-3,4-bisphosphate binding /  skeletal system morphogenesis / face morphogenesis / estrogen metabolic process / roof of mouth development ... skeletal system morphogenesis / face morphogenesis / estrogen metabolic process / roof of mouth development ... : / : /  luteinization / Leydig cell differentiation / ruffle organization / phosphatidylinositol biosynthetic process / phosphatidylinositol-3,4-bisphosphate binding / luteinization / Leydig cell differentiation / ruffle organization / phosphatidylinositol biosynthetic process / phosphatidylinositol-3,4-bisphosphate binding /  skeletal system morphogenesis / face morphogenesis / estrogen metabolic process / roof of mouth development / Synthesis of PIPs at the plasma membrane / platelet-derived growth factor receptor signaling pathway / androgen metabolic process / negative regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / post-embryonic development / skeletal system morphogenesis / face morphogenesis / estrogen metabolic process / roof of mouth development / Synthesis of PIPs at the plasma membrane / platelet-derived growth factor receptor signaling pathway / androgen metabolic process / negative regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / post-embryonic development /  PDZ domain binding / B cell receptor signaling pathway / establishment of protein localization / multicellular organism growth / ruffle membrane / cellular response to hydrogen peroxide / PDZ domain binding / B cell receptor signaling pathway / establishment of protein localization / multicellular organism growth / ruffle membrane / cellular response to hydrogen peroxide /  spermatogenesis / extracellular exosome / spermatogenesis / extracellular exosome /  nucleoplasm / nucleoplasm /  membrane / membrane /  plasma membrane / plasma membrane /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   HOMO SAPIENS (human) HOMO SAPIENS (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.4 Å MOLECULAR REPLACEMENT / Resolution: 1.4 Å | ||||||
 Authors Authors | Thomas, C.C. / Dowler, S. / Deak, M. / Alessi, D.R. / Van Aalten, D.M.F. | ||||||
 Citation Citation |  Journal: Biochem.J. / Year: 2001 Journal: Biochem.J. / Year: 2001Title: Crystal Structure of the Phosphatidylinositol 3,4-Bisphosphate-Binding Pleckstrin Homology (Ph) Domain of Tandem Ph-Domain-Containing Protein 1 (Tapp1): Molecular Basis of Lipid Specificity Authors: Thomas, C.C. / Dowler, S. / Deak, M. / Alessi, D.R. / Van Aalten, D.M.F. #1: Journal: Biochem.J. / Year: 2000 Title: Identification of Pleckstrin-Homology-Domain-Containing Proteins with Novel Phosphoinositide-Binding Specificities Authors: Dowler, S. / Currie, R.A. / Campbell, D.G. / Deak, M. / Kular, G. / Downes, C.P. / Alessi, D.R. #2:  Journal: Mol.Cell / Year: 2000 Journal: Mol.Cell / Year: 2000Title: Structural Basis for Discrimination of 3-Phosphoinositides by Pleckstrin Homology Domains Authors: Ferguson, K.M. / Kavran, J.M. / Sankaran, V.G. / Fournier, E. / Isakoff, S.J. / Skolnik, E.Y. / Lemmon, M.A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1eaz.cif.gz 1eaz.cif.gz | 65.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1eaz.ent.gz pdb1eaz.ent.gz | 47.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1eaz.json.gz 1eaz.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ea/1eaz https://data.pdbj.org/pub/pdb/validation_reports/ea/1eaz ftp://data.pdbj.org/pub/pdb/validation_reports/ea/1eaz ftp://data.pdbj.org/pub/pdb/validation_reports/ea/1eaz | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1fb8S S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 14240.303 Da / Num. of mol.: 1 / Fragment: PLECKSTRIN HOMOLOGY DOMAIN RESIDUES 182-303 Source method: isolated from a genetically manipulated source Details: ORDERED CITRATE MOLECULE IN LIPID BINDING POCKET / Source: (gene. exp.)   HOMO SAPIENS (human) / Plasmid: PGEX / Production host: HOMO SAPIENS (human) / Plasmid: PGEX / Production host:   ESCHERICHIA COLI (E. coli) / Strain (production host): BL21 / References: UniProt: Q9HB21 ESCHERICHIA COLI (E. coli) / Strain (production host): BL21 / References: UniProt: Q9HB21 |
|---|---|
| #2: Chemical | ChemComp-CIT /  Citric acid Citric acid |
| #3: Water | ChemComp-HOH /  Water Water |
| Sequence details | PROTEIN IS PH DOMAIN FROM ACCESSION NUMBER PROTEIN |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.48 Å3/Da / Density % sol: 48 % | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | pH: 5.6 Details: 0.085M SODIUM CITRATE, 25.5% PEG 4000, 15% GLYCEROL, 0.17M AMMONIUM ACETATE, pH 5.60 | ||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 20 ℃ / Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID14-4 / Wavelength: 0.9999 / Beamline: ID14-4 / Wavelength: 0.9999 |
| Detector | Type: ADSC CCD / Detector: CCD / Date: Feb 25, 2001 / Details: MIRRORS |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.9999 Å / Relative weight: 1 : 0.9999 Å / Relative weight: 1 |
| Reflection | Resolution: 1.4→27.524 Å / Num. obs: 27661 / % possible obs: 98.9 % / Observed criterion σ(I): 2 / Redundancy: 4 % / Rmerge(I) obs: 0.058 / Net I/σ(I): 21.7 |
| Reflection shell | Resolution: 1.4→1.45 Å / Redundancy: 3.1 % / Rmerge(I) obs: 0.115 / Mean I/σ(I) obs: 4.2 / % possible all: 97.7 |
| Reflection | *PLUS Lowest resolution: 30 Å |
| Reflection shell | *PLUS % possible obs: 97.7 % |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1FB8 Resolution: 1.4→30 Å / Cross valid method: THROUGHOUT / σ(F): 0 Details: MODEL BUILT WITH WARPNTRACE, REFINEMENT STARTED USING CNS THEN USED SHELXL USED ANISOTROPIC B-FACTORS IN SHELXL REFINEMENT.
| |||||||||||||||||||||||||||||||||
| Refine analyze | Occupancy sum non hydrogen: 990 | |||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.4→30 Å
| |||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||
| Software | *PLUS Name: SHELXL / Version: 97 / Classification: refinement | |||||||||||||||||||||||||||||||||
| Refinement | *PLUS Lowest resolution: 30 Å / % reflection Rfree: 5 % / Rfactor Rwork : 0.171 : 0.171 | |||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller











 PDBj
PDBj



