+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1o7b | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
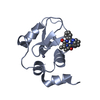
| Title | Refined solution structure of the human TSG-6 Link module | |||||||||
 Components Components | TUMOR NECROSIS FACTOR-INDUCIBLE PROTEIN TSG-6 | |||||||||
 Keywords Keywords |  CELL ADHESION / HYALURONAN-BINDING DOMAIN / CARBOHYDRATE-BINDING DOMAIN / CELL ADHESION / HYALURONAN-BINDING DOMAIN / CARBOHYDRATE-BINDING DOMAIN /  LINK MODULE / LINK MODULE /  GLYCOPROTEIN GLYCOPROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationfibronectin fibril organization / hyaluronan metabolic process / ovarian cumulus expansion /  hyaluronic acid binding / hyaluronic acid binding /  carboxylesterase activity / negative regulation of neutrophil chemotaxis / carboxylesterase activity / negative regulation of neutrophil chemotaxis /  Hydrolases; Acting on ester bonds; Carboxylic-ester hydrolases / negative regulation of osteoclast differentiation / Hydrolases; Acting on ester bonds; Carboxylic-ester hydrolases / negative regulation of osteoclast differentiation /  fibronectin binding / negative regulation of BMP signaling pathway ...fibronectin fibril organization / hyaluronan metabolic process / ovarian cumulus expansion / fibronectin binding / negative regulation of BMP signaling pathway ...fibronectin fibril organization / hyaluronan metabolic process / ovarian cumulus expansion /  hyaluronic acid binding / hyaluronic acid binding /  carboxylesterase activity / negative regulation of neutrophil chemotaxis / carboxylesterase activity / negative regulation of neutrophil chemotaxis /  Hydrolases; Acting on ester bonds; Carboxylic-ester hydrolases / negative regulation of osteoclast differentiation / Hydrolases; Acting on ester bonds; Carboxylic-ester hydrolases / negative regulation of osteoclast differentiation /  fibronectin binding / negative regulation of BMP signaling pathway / negative regulation of osteoblast differentiation / positive regulation of receptor clustering / negative regulation of inflammatory response / tertiary granule lumen / cell-cell signaling / ficolin-1-rich granule lumen / fibronectin binding / negative regulation of BMP signaling pathway / negative regulation of osteoblast differentiation / positive regulation of receptor clustering / negative regulation of inflammatory response / tertiary granule lumen / cell-cell signaling / ficolin-1-rich granule lumen /  cell adhesion / positive regulation of cell migration / cell adhesion / positive regulation of cell migration /  inflammatory response / inflammatory response /  calcium ion binding / Neutrophil degranulation / calcium ion binding / Neutrophil degranulation /  signal transduction / signal transduction /  extracellular space / extracellular region extracellular space / extracellular regionSimilarity search - Function | |||||||||
| Biological species |   HOMO SAPIENS (human) HOMO SAPIENS (human) | |||||||||
| Method |  SOLUTION NMR / AB INITIO SIMULATED ANNEALING SOLUTION NMR / AB INITIO SIMULATED ANNEALING | |||||||||
 Authors Authors | Blundell, C.D. / Teriete, P. / Kahmann, J.D. / Pickford, A.R. / Campbell, I.D. / Day, A.J. | |||||||||
 Citation Citation |  Journal: J. Biol. Chem. / Year: 2003 Journal: J. Biol. Chem. / Year: 2003Title: The link module from ovulation- and inflammation-associated protein TSG-6 changes conformation on hyaluronan binding. Authors: Blundell, C.D. / Mahoney, D.J. / Almond, A. / DeAngelis, P.L. / Kahmann, J.D. / Teriete, P. / Pickford, A.R. / Campbell, I.D. / Day, A.J. #1: Journal: J.Biol.Chem. / Year: 2002 Title: The Link Module from Human Tsg-6 Inhibits Neutrophil Migration in a Hyaluronan- and Inter-Alpha -Inhibitor-Independent Manner Authors: Getting, S.J. / Mahoney, D.J. / Cao, T. / Rugg, M.S. / Fries, E. / Milner, C.M. / Perretti, M. / Day, A.J. #2: Journal: J.Biol.Chem. / Year: 2002 Title: Hyaluronan Binding Properties of a Cd44 Chimera Containing the Link Module of Tsg-6 Authors: Lesley, J. / English, N.M. / Gal, I. / Mikecz, K. / Day, A.J. / Hyman, R. #3: Journal: J.Biol.Chem. / Year: 2001 Title: Mapping the Hyaluronan-Binding Site on the Link Module from Human Tumor Necrosis Factor-Stimulated Gene-6 by Site-Directed Mutagenesis Authors: Mahoney, D.J. / Blundell, C.D. / Day, A.J. #4: Journal: Structure / Year: 2000 Title: Localization and Characterization of the Hyaluronan-Binding Site on the Link Module from Human Tsg-6 Authors: Kahmann, J.D. / O'Brien, R. / Werner, J.M. / Heinegard, D. / Ladbury, J.E. / Campbell, I.D. / Day, A.J. #5: Journal: Protein Expr.Purif. / Year: 1997 Title: Method for Quantitative Refolding of the Link Module from Human Tsg-6 Authors: Kahmann, J.D. / Koruth, R. / Day, A.J. #6: Journal: Protein Expr.Purif. / Year: 1996 Title: Overexpression, Purification, and Refolding of Link Module from Human Tsg-6 in Escherichia Coli: Effect of Temperature, Media, and Mutagenesis on Lysine Misincorporation at Arginine Aga Codons Authors: Day, A.J. / Aplin, R.T. / Willis, A.C. #7: Journal: Cell(Cambridge,Mass.) / Year: 1996 Title: Solution Structure of the Link Module: A Hyaluronan-Binding Domain Involved in Extracellular Matrix Stability and Cell Migration Authors: Kohda, D. / Morton, C.J. / Parkar, A.A. / Hatanaka, H. / Inagaki, F.M. / Campbell, I.D. / Day, A.J. #8: Journal: J.Cell Biol. / Year: 1992 Title: A Novel Secretory Tumour Necrosis Factor-Inducible Protein (Tsg-6) is a Member of the Family of Hyaluronate Binding Proteins, Closely Related to the Adhesion Receptor Cd44 Authors: Lee, T.H. / Wisniewski, H.G. / Vilcek, J. | |||||||||
| History |
| |||||||||
| Remark 700 | SHEET THE SHEET STRUCTURE OF THIS MOLECULE IS BIFURCATED. IN ORDER TO REPRESENT THIS FEATURE IN ... SHEET THE SHEET STRUCTURE OF THIS MOLECULE IS BIFURCATED. IN ORDER TO REPRESENT THIS FEATURE IN THE SHEET RECORDS BELOW, TWO SHEETS ARE DEFINED. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1o7b.cif.gz 1o7b.cif.gz | 593.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1o7b.ent.gz pdb1o7b.ent.gz | 516.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1o7b.json.gz 1o7b.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/o7/1o7b https://data.pdbj.org/pub/pdb/validation_reports/o7/1o7b ftp://data.pdbj.org/pub/pdb/validation_reports/o7/1o7b ftp://data.pdbj.org/pub/pdb/validation_reports/o7/1o7b | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 10946.578 Da / Num. of mol.: 1 / Fragment: LINK_MODULE, RESIDUES 36-133 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   HOMO SAPIENS (human) / Description: EXTRACELLULAR, INFLAMMATION-ASSOCIATED / Plasmid: PRK172 / Production host: HOMO SAPIENS (human) / Description: EXTRACELLULAR, INFLAMMATION-ASSOCIATED / Plasmid: PRK172 / Production host:   ESCHERICHIA COLI (E. coli) / Strain (production host): BL21(DE3) / References: UniProt: P98066 ESCHERICHIA COLI (E. coli) / Strain (production host): BL21(DE3) / References: UniProt: P98066 |
|---|---|
| Compound details | POSSIBLE ROLE IN CELL-MATRIX INTERACTIO |
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||
| NMR details | Text: STRUCTURE DETERMINED BY NMR SPECTROSCOPY ON UNIFORMLY 15N- AND 13C,15N-LABELLED LINK_TSG6 |
- Sample preparation
Sample preparation
| Details | Contents: 10% D2O, 1-3MM LINK_TSG6 |
|---|---|
| Sample conditions | Ionic strength: ~2MM NA IONS / pH: 6.0 / Pressure: 1 atm / Temperature: 298 K |
Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: AB INITIO SIMULATED ANNEALING / Software ordinal: 1 Details: REFINEMENT DETAILS CAN BE FOUND IN THE JRNL CITATION ABOVE | ||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: LEAST RESTRAINT VIOLATION / Conformers calculated total number: 250 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller













 PDBj
PDBj




