+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | E. coli clamp loader with closed clamp on primed template DNA | |||||||||
 Map data Map data | CLC.DNA.Beta2 Closed | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | clamp loader /  DNA clamp / DNA clamp /  AAA+ / AAA+ /  ATPase / DNA BINDING PROTEIN-DNA complex ATPase / DNA BINDING PROTEIN-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology information DNA polymerase III, clamp loader complex / Hda-beta clamp complex / bacterial-type DNA replication / replication inhibiting complex / DNA polymerase III, clamp loader complex / Hda-beta clamp complex / bacterial-type DNA replication / replication inhibiting complex /  DNA polymerase III complex / DNA polymerase III complex /  replisome / regulation of DNA-templated DNA replication initiation / DNA strand elongation involved in DNA replication / DNA polymerase processivity factor activity / error-prone translesion synthesis ... replisome / regulation of DNA-templated DNA replication initiation / DNA strand elongation involved in DNA replication / DNA polymerase processivity factor activity / error-prone translesion synthesis ... DNA polymerase III, clamp loader complex / Hda-beta clamp complex / bacterial-type DNA replication / replication inhibiting complex / DNA polymerase III, clamp loader complex / Hda-beta clamp complex / bacterial-type DNA replication / replication inhibiting complex /  DNA polymerase III complex / DNA polymerase III complex /  replisome / regulation of DNA-templated DNA replication initiation / DNA strand elongation involved in DNA replication / DNA polymerase processivity factor activity / error-prone translesion synthesis / negative regulation of DNA-templated DNA replication initiation / 3'-5' exonuclease activity / ribonucleoside triphosphate phosphatase activity / response to radiation / DNA-templated DNA replication / replisome / regulation of DNA-templated DNA replication initiation / DNA strand elongation involved in DNA replication / DNA polymerase processivity factor activity / error-prone translesion synthesis / negative regulation of DNA-templated DNA replication initiation / 3'-5' exonuclease activity / ribonucleoside triphosphate phosphatase activity / response to radiation / DNA-templated DNA replication /  DNA replication / DNA replication /  DNA-directed DNA polymerase / DNA-directed DNA polymerase /  DNA-directed DNA polymerase activity / DNA damage response / DNA-directed DNA polymerase activity / DNA damage response /  ATP hydrolysis activity / protein homodimerization activity / ATP hydrolysis activity / protein homodimerization activity /  DNA binding / DNA binding /  ATP binding / identical protein binding / ATP binding / identical protein binding /  cytosol cytosolSimilarity search - Function | |||||||||
| Biological species |   Escherichia coli (E. coli) / Escherichia coli (E. coli) /   Escherichia coli K-12 (bacteria) Escherichia coli K-12 (bacteria) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.6 Å cryo EM / Resolution: 2.6 Å | |||||||||
 Authors Authors | Oakley AJ / Xu Z-Q / Dixon NE | |||||||||
| Funding support |  Australia, 1 items Australia, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structural characterisation of the complete cycle of sliding clamp loading in E. coli Authors: Xu Z-Q / Oakley AJ / Dixon NE | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40083.map.gz emd_40083.map.gz | 140.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40083-v30.xml emd-40083-v30.xml emd-40083.xml emd-40083.xml | 29.7 KB 29.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_40083.png emd_40083.png | 76 KB | ||
| Filedesc metadata |  emd-40083.cif.gz emd-40083.cif.gz | 8.6 KB | ||
| Others |  emd_40083_half_map_1.map.gz emd_40083_half_map_1.map.gz emd_40083_half_map_2.map.gz emd_40083_half_map_2.map.gz | 140.6 MB 140.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40083 http://ftp.pdbj.org/pub/emdb/structures/EMD-40083 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40083 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40083 | HTTPS FTP |
-Related structure data
| Related structure data |  8gj2MC  8giyC  8gizC  8gj0C  8gj1C  8gj3C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40083.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40083.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CLC.DNA.Beta2 Closed | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.66 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: CLC.DNA.Beta2 Closed half map 2
| File | emd_40083_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CLC.DNA.Beta2 Closed half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
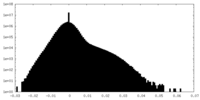
| Density Histograms |
-Half map: CLC.DNA.Beta2 Closed half map 1
| File | emd_40083_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CLC.DNA.Beta2 Closed half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : E. coli clamp loader with closed clamp on primed template DNA
+Supramolecule #1: E. coli clamp loader with closed clamp on primed template DNA
+Macromolecule #1: DNA polymerase III subunit delta
+Macromolecule #2: DNA polymerase III subunit tau
+Macromolecule #3: DNA polymerase III subunit delta'
+Macromolecule #4: DNA polymerase III subunit psi
+Macromolecule #5: Beta sliding clamp
+Macromolecule #6: Primer
+Macromolecule #7: Template
+Macromolecule #8: ADENOSINE-5'-DIPHOSPHATE
+Macromolecule #9: MAGNESIUM ION
+Macromolecule #10: TETRAFLUOROALUMINATE ION
+Macromolecule #11: ZINC ION
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.5 mg/mL | |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.6 Component:
Details: 30 mM Tris-HCl pH 7.6, 5 mM MgCl2, 2 mM ADP, 0.5 mM AlCl3, 5 mM NaF, 5 mM dithiothreitol, 0.25 mM EDTA, 2% glycerol. | |||||||||||||||||||||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 120 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.03 kPa Details: Used a Zepto Low-pressure plasma systems (PLASMA CLEANER) (Diener Electronic) at 10% power for 120 seconds. | |||||||||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 279 K / Instrument: FEI VITROBOT MARK IV Details: 3 uL of sample was applied onto a Ultrafoil Au R1.2/1.3 grid. Blot for 4.5 s with no extra force before plunging into liquid ethane.. | |||||||||||||||||||||||||||
| Details | 30 microL of 6.0 microM delta/tau3/delta' complex was mixed with chi/psi complex, beta, and p/t DNA (annealed DNA oligonucleotides) at a molar ratio of 1:1.3 and dialysed at 4 degrees C against 250 mL of 30 mM Tris-HCl pH 7.6, 5 mM MgCl2, 2 mM ADP, 0.5 mM AlCl3, 5 mM NaF, 5 mM dithiothreitol, 0.25 mM EDTA, 2% glycerol. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 1.3 µm / Nominal defocus min: 0.4 µm / Nominal magnification: 165000 Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 1.3 µm / Nominal defocus min: 0.4 µm / Nominal magnification: 165000 |
| Specialist optics | Energy filter - Name: GIF Bioquantum / Details: unfiltered mode |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Digitization - Frames/image: 1-60 / Number grids imaged: 1 / Number real images: 1703 / Average exposure time: 6.0 sec. / Average electron dose: 60.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Particle selection | Number selected: 600000 |
|---|---|
| Startup model | Type of model: OTHER / Details: Ab-initio model from RELION |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.1) |
| Final 3D classification | Number classes: 4 / Avg.num./class: 120000 / Software - Name: RELION (ver. 3.1) Details: Most particles were classified to two classes that were used for final 3D reconstruction. |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.1) |
| Final reconstruction | Number classes used: 2 / Applied symmetry - Point group: C1 (asymmetric) / Algorithm: BACK PROJECTION / Resolution.type: BY AUTHOR / Resolution: 2.6 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.1) / Number images used: 452000 |
 Movie
Movie Controller
Controller









 Z
Z Y
Y X
X