[English] 日本語
 Yorodumi
Yorodumi- EMDB-27757: Cryo-EM Structure of Eastern Equine Encephalitis Virus in complex... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
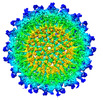
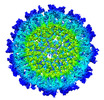
| Title | Cryo-EM Structure of Eastern Equine Encephalitis Virus in complex with SKE26 Fab | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  Eastern Equine Encephalitis virus / Eastern Equine Encephalitis virus /  Virus Like Particle / virus-fab complex Virus Like Particle / virus-fab complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationT=4 icosahedral viral capsid / host cell cytoplasm / symbiont entry into host cell / serine-type endopeptidase activity / fusion of virus membrane with host endosome membrane / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity /  proteolysis ...T=4 icosahedral viral capsid / host cell cytoplasm / symbiont entry into host cell / serine-type endopeptidase activity / fusion of virus membrane with host endosome membrane / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity / proteolysis ...T=4 icosahedral viral capsid / host cell cytoplasm / symbiont entry into host cell / serine-type endopeptidase activity / fusion of virus membrane with host endosome membrane / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity /  proteolysis / proteolysis /  plasma membrane / plasma membrane /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |    Eastern equine encephalitis virus / Eastern equine encephalitis virus /   Macaca fascicularis (crab-eating macaque) Macaca fascicularis (crab-eating macaque) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.5 Å cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Pletnev S / Verardi R / Roedeger M / Kwong P | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2023 Journal: Cell / Year: 2023Title: Vaccine elicitation and structural basis for antibody protection against alphaviruses. Authors: Matthew S Sutton / Sergei Pletnev / Victoria Callahan / Sungyoul Ko / Yaroslav Tsybovsky / Tatsiana Bylund / Ryan G Casner / Gabriele Cerutti / Christina L Gardner / Veronica Guirguis / ...Authors: Matthew S Sutton / Sergei Pletnev / Victoria Callahan / Sungyoul Ko / Yaroslav Tsybovsky / Tatsiana Bylund / Ryan G Casner / Gabriele Cerutti / Christina L Gardner / Veronica Guirguis / Raffaello Verardi / Baoshan Zhang / David Ambrozak / Margaret Beddall / Hong Lei / Eun Sung Yang / Tracy Liu / Amy R Henry / Reda Rawi / Arne Schön / Chaim A Schramm / Chen-Hsiang Shen / Wei Shi / Tyler Stephens / Yongping Yang / Maria Burgos Florez / Julie E Ledgerwood / Crystal W Burke / Lawrence Shapiro / Julie M Fox / Peter D Kwong / Mario Roederer /  Abstract: Alphaviruses are RNA viruses that represent emerging public health threats. To identify protective antibodies, we immunized macaques with a mixture of western, eastern, and Venezuelan equine ...Alphaviruses are RNA viruses that represent emerging public health threats. To identify protective antibodies, we immunized macaques with a mixture of western, eastern, and Venezuelan equine encephalitis virus-like particles (VLPs), a regimen that protects against aerosol challenge with all three viruses. Single- and triple-virus-specific antibodies were isolated, and we identified 21 unique binding groups. Cryo-EM structures revealed that broad VLP binding inversely correlated with sequence and conformational variability. One triple-specific antibody, SKT05, bound proximal to the fusion peptide and neutralized all three Env-pseudotyped encephalitic alphaviruses by using different symmetry elements for recognition across VLPs. Neutralization in other assays (e.g., chimeric Sindbis virus) yielded variable results. SKT05 bound backbone atoms of sequence-diverse residues, enabling broad recognition despite sequence variability; accordingly, SKT05 protected mice against Venezuelan equine encephalitis virus, chikungunya virus, and Ross River virus challenges. Thus, a single vaccine-elicited antibody can protect in vivo against a broad range of alphaviruses. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27757.map.gz emd_27757.map.gz | 1.2 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27757-v30.xml emd-27757-v30.xml emd-27757.xml emd-27757.xml | 21 KB 21 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27757_fsc.xml emd_27757_fsc.xml | 22.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_27757.png emd_27757.png | 58.6 KB | ||
| Masks |  emd_27757_msk_1.map emd_27757_msk_1.map | 1.3 GB |  Mask map Mask map | |
| Others |  emd_27757_half_map_1.map.gz emd_27757_half_map_1.map.gz emd_27757_half_map_2.map.gz emd_27757_half_map_2.map.gz | 1.2 GB 1.2 GB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27757 http://ftp.pdbj.org/pub/emdb/structures/EMD-27757 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27757 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27757 | HTTPS FTP |
-Related structure data
| Related structure data |  8dwoMC  8decC  8dedC  8deeC  8defC  8deqC  8derC  8dulC  8dunC  8eeuC  8eevC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27757.map.gz / Format: CCP4 / Size: 1.3 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27757.map.gz / Format: CCP4 / Size: 1.3 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.66 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_27757_msk_1.map emd_27757_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_27757_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_27757_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Eastern equine encephalitis virus
| Entire | Name:    Eastern equine encephalitis virus Eastern equine encephalitis virus |
|---|---|
| Components |
|
-Supramolecule #1: Eastern equine encephalitis virus
| Supramolecule | Name: Eastern equine encephalitis virus / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 11021 / Sci species name: Eastern equine encephalitis virus / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: OTHER / Virus enveloped: Yes / Virus empty: Yes |
|---|
-Macromolecule #1: Envelope glycoprotein E1
| Macromolecule | Name: Envelope glycoprotein E1 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO / EC number:  togavirin togavirin |
|---|---|
| Source (natural) | Organism:    Eastern equine encephalitis virus Eastern equine encephalitis virus |
| Molecular weight | Theoretical: 47.961215 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: YEHTAVMPNK VGIPYKALVE RPGYAPVHLQ IQLVNTRIIP STNLEYITCK YKTKVPSPVV KCCGATQCTS KPHPDYQCQV FSGVYPFMY GGAYCFCDTE NTQMSEAYVE RSEECSIDHA KAYKVHTGTV QAMVNITYGS VSWRSADVYV NGETPAKIGD A KLIIGPLS ...String: YEHTAVMPNK VGIPYKALVE RPGYAPVHLQ IQLVNTRIIP STNLEYITCK YKTKVPSPVV KCCGATQCTS KPHPDYQCQV FSGVYPFMY GGAYCFCDTE NTQMSEAYVE RSEECSIDHA KAYKVHTGTV QAMVNITYGS VSWRSADVYV NGETPAKIGD A KLIIGPLS SAWSPFDNKV VVYGHEVYNY DFPEYGTGKA GSFGDLQSRT STSNDLYANT NLKLQRPQAG IVHTPFTQVP SG FERWKKD KGAPLNDVAP FGCSIALEPL RAENCAVGSI PISIDIPDAA FTRISETPTV SDLECKITEC TYAFDFGGIA TVA YKSSKA GNCPIHSPSG VAVIKENDVT LAESGSFTFH FSTANIHPAF KLQVCTSAVT CKGDCKPPKD HIVDYPAQHT ESFT SAISA TAWSWIKVLV GGTSAFIVLG LIATAVVALV LFFHRH UniProtKB: Structural polyprotein |
-Macromolecule #2: Envelope glycoprotein E2
| Macromolecule | Name: Envelope glycoprotein E2 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO / EC number:  togavirin togavirin |
|---|---|
| Source (natural) | Organism:    Eastern equine encephalitis virus Eastern equine encephalitis virus |
| Molecular weight | Theoretical: 47.04702 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DLDTHFTQYK LARPYIADCP NCGHSRCDSP IAIEEVRGDA HAGVIRIQTS AMFGLKTDGV DLAYMSFMNG KTQKSIKIDN LHVRTSAPC SLVSHHGYYI LAQCPPGDTV TVGFHDGPNR HTCTVAHKVE FRPVGREKYR HPPEHGVELP CNRYTHKRAD Q GHYVEMHQ ...String: DLDTHFTQYK LARPYIADCP NCGHSRCDSP IAIEEVRGDA HAGVIRIQTS AMFGLKTDGV DLAYMSFMNG KTQKSIKIDN LHVRTSAPC SLVSHHGYYI LAQCPPGDTV TVGFHDGPNR HTCTVAHKVE FRPVGREKYR HPPEHGVELP CNRYTHKRAD Q GHYVEMHQ PGLVADHSLL SIHSAKVKIT VPSGAQVKYY CKCPDVRKGI TSSDHTTTCT DVKQCRAYLI DNKKWVYNSG RL PRGEGDT FKGKLHVPFV PVKAKCIATL APEPLVEHKH RTLILHLHPD HPTLLTTRSL GSDANPTRQW IERPTTVNFT VTG EGLEYT WGNHPPKRVW AQESGEGNPH GWPHEVVVYY YNRYPLTTII GLCTCVAIIM VSCVTSVWLL CRTRNLCITP YKLA PNAQV PILLALLCCI KPTRA UniProtKB: Structural polyprotein |
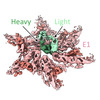
-Macromolecule #3: SKE26 Fab Heavy Chain
| Macromolecule | Name: SKE26 Fab Heavy Chain / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Macaca fascicularis (crab-eating macaque) Macaca fascicularis (crab-eating macaque) |
| Molecular weight | Theoretical: 25.358479 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: VVQLVQSGAE VKRPGESLRI SCKTSGYRFT DIWINWVRQR PGKGLEWMGS IYLDDSDIRF NPSFQGHVTI SADKSTSTTY LQWSSLKAS DTATYFCARS VWSGYYIGWF DVWGPGVLVT VSSASTKGPS VFPLAPSSRS TSESTAALGC LVKDYFPEPV T VSWNSGSL ...String: VVQLVQSGAE VKRPGESLRI SCKTSGYRFT DIWINWVRQR PGKGLEWMGS IYLDDSDIRF NPSFQGHVTI SADKSTSTTY LQWSSLKAS DTATYFCARS VWSGYYIGWF DVWGPGVLVT VSSASTKGPS VFPLAPSSRS TSESTAALGC LVKDYFPEPV T VSWNSGSL TSGVHTFPAV LQSSGLYSLS SVVTVPSSSL GTQTYVCNVN HKPSNTKVDK RVEIKTCGGL EVLFQ |
-Macromolecule #4: SKE26 Fab Light Chain
| Macromolecule | Name: SKE26 Fab Light Chain / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Macaca fascicularis (crab-eating macaque) Macaca fascicularis (crab-eating macaque) |
| Molecular weight | Theoretical: 23.542113 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DVVMTQTPLS LPVTPGEPAS ISCRSSQSLL DSNGNTYLHW YLQKPGQSPQ LLIYGGSNRA SGVPDRFSGS GSGTDFTLTI NNVEAGDVG TYYCMQVIQV PLTFGGGTKV DIKRTVAAPS VFIFPPSEDQ VKSGTVSVVC LLNNFYPREA SVKWKVDGAL K TGNSQESV ...String: DVVMTQTPLS LPVTPGEPAS ISCRSSQSLL DSNGNTYLHW YLQKPGQSPQ LLIYGGSNRA SGVPDRFSGS GSGTDFTLTI NNVEAGDVG TYYCMQVIQV PLTFGGGTKV DIKRTVAAPS VFIFPPSEDQ VKSGTVSVVC LLNNFYPREA SVKWKVDGAL K TGNSQESV TEQDSKDNTY SLSSTLTLSS TEYQSHKVYA CEVTHQGLSS PVTKSFNRGE C |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.7000000000000001 µm Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.7000000000000001 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 5662 / Average electron dose: 58.06 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





















 Z
Z Y
Y X
X



























