[English] 日本語
 Yorodumi
Yorodumi- EMDB-21302: Human Diacylglycerol Acyltransferase 1 in complex with oleoyl-CoA -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21302 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human Diacylglycerol Acyltransferase 1 in complex with oleoyl-CoA Diglyceride acyltransferase Diglyceride acyltransferase | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords |  diacylglycerol acyltransferase 1 / diacylglycerol acyltransferase 1 /  MBOAT / oleoyl-CoA / ER / MBOAT / oleoyl-CoA / ER /  MEMBRANE PROTEIN MEMBRANE PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology information retinol O-fatty-acyltransferase / retinol O-fatty-acyltransferase /  2-acylglycerol O-acyltransferase activity / 2-acylglycerol O-acyltransferase activity /  retinol O-fatty-acyltransferase activity / monoacylglycerol biosynthetic process / Triglyceride biosynthesis / diacylglycerol metabolic process / Acyl chain remodeling of DAG and TAG / triglyceride biosynthetic process / long-chain fatty-acyl-CoA metabolic process / retinol O-fatty-acyltransferase activity / monoacylglycerol biosynthetic process / Triglyceride biosynthesis / diacylglycerol metabolic process / Acyl chain remodeling of DAG and TAG / triglyceride biosynthetic process / long-chain fatty-acyl-CoA metabolic process /  diacylglycerol O-acyltransferase ... diacylglycerol O-acyltransferase ... retinol O-fatty-acyltransferase / retinol O-fatty-acyltransferase /  2-acylglycerol O-acyltransferase activity / 2-acylglycerol O-acyltransferase activity /  retinol O-fatty-acyltransferase activity / monoacylglycerol biosynthetic process / Triglyceride biosynthesis / diacylglycerol metabolic process / Acyl chain remodeling of DAG and TAG / triglyceride biosynthetic process / long-chain fatty-acyl-CoA metabolic process / retinol O-fatty-acyltransferase activity / monoacylglycerol biosynthetic process / Triglyceride biosynthesis / diacylglycerol metabolic process / Acyl chain remodeling of DAG and TAG / triglyceride biosynthetic process / long-chain fatty-acyl-CoA metabolic process /  diacylglycerol O-acyltransferase / diacylglycerol O-acyltransferase /  diacylglycerol O-acyltransferase activity / very-low-density lipoprotein particle assembly / diacylglycerol O-acyltransferase activity / very-low-density lipoprotein particle assembly /  lipid storage / triglyceride metabolic process / lipid storage / triglyceride metabolic process /  acyltransferase activity / fatty acid homeostasis / specific granule membrane / Neutrophil degranulation / endoplasmic reticulum membrane / acyltransferase activity / fatty acid homeostasis / specific granule membrane / Neutrophil degranulation / endoplasmic reticulum membrane /  membrane / identical protein binding / membrane / identical protein binding /  plasma membrane plasma membraneSimilarity search - Function | ||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.1 Å cryo EM / Resolution: 3.1 Å | ||||||||||||
 Authors Authors | Wang L / Qian H | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nature / Year: 2020 Journal: Nature / Year: 2020Title: Structure and mechanism of human diacylglycerol O-acyltransferase 1. Authors: Lie Wang / Hongwu Qian / Yin Nian / Yimo Han / Zhenning Ren / Hanzhi Zhang / Liya Hu / B V Venkataram Prasad / Arthur Laganowsky / Nieng Yan / Ming Zhou /   Abstract: Diacylglycerol O-acyltransferase 1 (DGAT1) synthesizes triacylglycerides and is required for dietary fat absorption and fat storage in humans. DGAT1 belongs to the membrane-bound O-acyltransferase ...Diacylglycerol O-acyltransferase 1 (DGAT1) synthesizes triacylglycerides and is required for dietary fat absorption and fat storage in humans. DGAT1 belongs to the membrane-bound O-acyltransferase (MBOAT) superfamily, members of which are found in all kingdoms of life and are involved in the acylation of lipids and proteins. How human DGAT1 and other mammalian members of the MBOAT family recognize their substrates and catalyse their reactions is unknown. The absence of three-dimensional structures also hampers rational targeting of DGAT1 for therapeutic purposes. Here we present the cryo-electron microscopy structure of human DGAT1 in complex with an oleoyl-CoA substrate. Each DGAT1 protomer has nine transmembrane helices, eight of which form a conserved structural fold that we name the MBOAT fold. The MBOAT fold in DGAT1 forms a hollow chamber in the membrane that encloses highly conserved catalytic residues. The chamber has separate entrances for each of the two substrates, fatty acyl-CoA and diacylglycerol. DGAT1 can exist as either a homodimer or a homotetramer and the two forms have similar enzymatic activity. The N terminus of DGAT1 interacts with the neighbouring protomer and these interactions are required for enzymatic activity. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21302.map.gz emd_21302.map.gz | 28.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21302-v30.xml emd-21302-v30.xml emd-21302.xml emd-21302.xml | 16.6 KB 16.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_21302_fsc.xml emd_21302_fsc.xml | 9.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_21302.png emd_21302.png | 266.9 KB | ||
| Filedesc metadata |  emd-21302.cif.gz emd-21302.cif.gz | 6.1 KB | ||
| Others |  emd_21302_half_map_1.map.gz emd_21302_half_map_1.map.gz emd_21302_half_map_2.map.gz emd_21302_half_map_2.map.gz | 28.3 MB 28.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21302 http://ftp.pdbj.org/pub/emdb/structures/EMD-21302 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21302 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21302 | HTTPS FTP |
-Related structure data
| Related structure data |  6vp0MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_21302.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21302.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.114 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: #1
| File | emd_21302_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
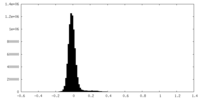
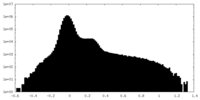
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_21302_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
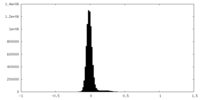
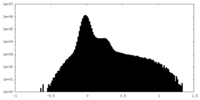
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : human diacylglycerol acyltransferase 1 in complex with oleoyl-CoA
| Entire | Name: human diacylglycerol acyltransferase 1 in complex with oleoyl-CoA Diglyceride acyltransferase Diglyceride acyltransferase |
|---|---|
| Components |
|
-Supramolecule #1: human diacylglycerol acyltransferase 1 in complex with oleoyl-CoA
| Supramolecule | Name: human diacylglycerol acyltransferase 1 in complex with oleoyl-CoA type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Diacylglycerol O-acyltransferase 1
| Macromolecule | Name: Diacylglycerol O-acyltransferase 1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number:  diacylglycerol O-acyltransferase diacylglycerol O-acyltransferase |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 55.339133 KDa |
| Recombinant expression | Organism:   Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MGDRGSSRRR RTGSRPSSHG GGGPAAAEEE VRDAAAGPDV GAAGDAPAPA PNKDGDAGVG SGHWELRCHR LQDSLFSSDS GFSNYRGIL NWCVVMLILS NARLFLENLI KYGILVDPIQ VVSLFLKDPY SWPAPCLVIA ANVFAVAAFQ VEKRLAVGAL T EQAGLLLH ...String: MGDRGSSRRR RTGSRPSSHG GGGPAAAEEE VRDAAAGPDV GAAGDAPAPA PNKDGDAGVG SGHWELRCHR LQDSLFSSDS GFSNYRGIL NWCVVMLILS NARLFLENLI KYGILVDPIQ VVSLFLKDPY SWPAPCLVIA ANVFAVAAFQ VEKRLAVGAL T EQAGLLLH VANLATILCF PAAVVLLVES ITPVGSLLAL MAHTILFLKL FSYRDVNSWC RRARAKAASA GKKASSAAAP HT VSYPDNL TYRDLYYFLF APTLCYELNF PRSPRIRKRF LLRRILEMLF FTQLQVGLIQ QWMVPTIQNS MKPFKDMDYS RII ERLLKL AVPNHLIWLI FFYWLFHSCL NAVAELMQFG DREFYRDWWN SESVTYFWQN WNIPVHKWCI RHFYKPMLRR GSSK WMART GVFLASAFFH EYLVSVPLRM FRLWAFTGMM AQIPLAWFVG RFFQGNYGNA AVWLSLIIGQ PIAVLMYVHD YYVLN YEAP AAEA UniProtKB:  Diacylglycerol O-acyltransferase 1 Diacylglycerol O-acyltransferase 1 |
-Macromolecule #2: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(tri...
| Macromolecule | Name: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate type: ligand / ID: 2 / Number of copies: 4 / Formula: POV |
|---|---|
| Molecular weight | Theoretical: 760.076 Da |
| Chemical component information |  ChemComp-POV: |
-Macromolecule #3: O-[(R)-{[(2R)-2,3-bis(octadecanoyloxy)propyl]oxy}(hydroxy)phospho...
| Macromolecule | Name: O-[(R)-{[(2R)-2,3-bis(octadecanoyloxy)propyl]oxy}(hydroxy)phosphoryl]-L-serine type: ligand / ID: 3 / Number of copies: 4 / Formula: P5S |
|---|---|
| Molecular weight | Theoretical: 792.075 Da |
| Chemical component information |  ChemComp-P5S: |
-Macromolecule #4: S-{(3R,5R,9R)-1-[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-4-hydrox...
| Macromolecule | Name: S-{(3R,5R,9R)-1-[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-4-hydroxy-3-(phosphonooxy)tetrahydrofuran-2-yl]-3,5,9-trihydroxy-8,8-dimethyl-3,5-dioxido-10,14-dioxo-2,4,6-trioxa-11,15-diaza- ...Name: S-{(3R,5R,9R)-1-[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-4-hydroxy-3-(phosphonooxy)tetrahydrofuran-2-yl]-3,5,9-trihydroxy-8,8-dimethyl-3,5-dioxido-10,14-dioxo-2,4,6-trioxa-11,15-diaza-3lambda~5~,5lambda~5~-diphosphaheptadecan-17-yl} (9Z)-octadec-9-enethioate (non-preferred name) type: ligand / ID: 4 / Number of copies: 2 / Formula: 3VV |
|---|---|
| Molecular weight | Theoretical: 1.03198 KDa |
| Chemical component information |  ChemComp-3VV: |
-Macromolecule #5: Lauryl Maltose Neopentyl Glycol
| Macromolecule | Name: Lauryl Maltose Neopentyl Glycol / type: ligand / ID: 5 / Number of copies: 2 / Formula: AV0 |
|---|---|
| Molecular weight | Theoretical: 1.005188 KDa |
| Chemical component information |  ChemComp-AV0: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 20 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 305 K / Instrument: FEI VITROBOT MARK IV |
| Details | This sample was monodisperse. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 50.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)