+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cysteine tRNA ligase homodimer | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  cysteine / cysteine /  trna / trna /  ligase / synthetase / ligase / synthetase /  homodimer / homodimer /  RNA RNA | |||||||||
| Function / homology |  Function and homology information Function and homology information cysteine-tRNA ligase / cysteine-tRNA ligase /  cysteine-tRNA ligase activity / cysteinyl-tRNA aminoacylation / zinc ion binding / cysteine-tRNA ligase activity / cysteinyl-tRNA aminoacylation / zinc ion binding /  ATP binding / ATP binding /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |    Pyrococcus furiosus DSM 3638 (archaea) Pyrococcus furiosus DSM 3638 (archaea) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.8 Å cryo EM / Resolution: 2.8 Å | |||||||||
 Authors Authors | Pacesa M / Correia BE / Levy ED | |||||||||
| Funding support | European Union, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2024 Journal: Cell / Year: 2024Title: An atlas of protein homo-oligomerization across domains of life. Authors: Hugo Schweke / Martin Pacesa / Tal Levin / Casper A Goverde / Prasun Kumar / Yoan Duhoo / Lars J Dornfeld / Benjamin Dubreuil / Sandrine Georgeon / Sergey Ovchinnikov / Derek N Woolfson / ...Authors: Hugo Schweke / Martin Pacesa / Tal Levin / Casper A Goverde / Prasun Kumar / Yoan Duhoo / Lars J Dornfeld / Benjamin Dubreuil / Sandrine Georgeon / Sergey Ovchinnikov / Derek N Woolfson / Bruno E Correia / Sucharita Dey / Emmanuel D Levy /      Abstract: Protein structures are essential to understanding cellular processes in molecular detail. While advances in artificial intelligence revealed the tertiary structure of proteins at scale, their ...Protein structures are essential to understanding cellular processes in molecular detail. While advances in artificial intelligence revealed the tertiary structure of proteins at scale, their quaternary structure remains mostly unknown. We devise a scalable strategy based on AlphaFold2 to predict homo-oligomeric assemblies across four proteomes spanning the tree of life. Our results suggest that approximately 45% of an archaeal proteome and a bacterial proteome and 20% of two eukaryotic proteomes form homomers. Our predictions accurately capture protein homo-oligomerization, recapitulate megadalton complexes, and unveil hundreds of homo-oligomer types, including three confirmed experimentally by structure determination. Integrating these datasets with omics information suggests that a majority of known protein complexes are symmetric. Finally, these datasets provide a structural context for interpreting disease mutations and reveal coiled-coil regions as major enablers of quaternary structure evolution in human. Our strategy is applicable to any organism and provides a comprehensive view of homo-oligomerization in proteomes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18415.map.gz emd_18415.map.gz | 51.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18415-v30.xml emd-18415-v30.xml emd-18415.xml emd-18415.xml | 17 KB 17 KB | Display Display |  EMDB header EMDB header |
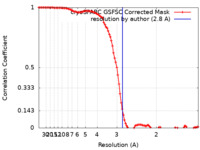
| FSC (resolution estimation) |  emd_18415_fsc.xml emd_18415_fsc.xml | 9.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_18415.png emd_18415.png | 65.9 KB | ||
| Filedesc metadata |  emd-18415.cif.gz emd-18415.cif.gz | 6 KB | ||
| Others |  emd_18415_half_map_1.map.gz emd_18415_half_map_1.map.gz emd_18415_half_map_2.map.gz emd_18415_half_map_2.map.gz | 95.5 MB 95.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18415 http://ftp.pdbj.org/pub/emdb/structures/EMD-18415 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18415 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18415 | HTTPS FTP |
-Related structure data
| Related structure data |  8qhpMC  8p49C  8q70C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_18415.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18415.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.726 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_18415_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
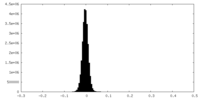
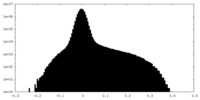
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_18415_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
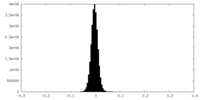
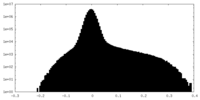
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Homodimeric form of Cysteine tRNA ligase
| Entire | Name: Homodimeric form of Cysteine tRNA ligase |
|---|---|
| Components |
|
-Supramolecule #1: Homodimeric form of Cysteine tRNA ligase
| Supramolecule | Name: Homodimeric form of Cysteine tRNA ligase / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:    Pyrococcus furiosus DSM 3638 (archaea) Pyrococcus furiosus DSM 3638 (archaea) |
| Molecular weight | Theoretical: 114 KDa |
-Macromolecule #1: Cysteine--tRNA ligase
| Macromolecule | Name: Cysteine--tRNA ligase / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:    Pyrococcus furiosus DSM 3638 (archaea) Pyrococcus furiosus DSM 3638 (archaea) |
| Molecular weight | Theoretical: 57.558605 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MKHHHHHHGG SSGGLKVYNT LTKQKEEFKP LREGEVKMYV CGPTVYDYPH LGHARTYIAF DVIRRYLEHK GYTVLMVMNF TDIDDKIIK RARETGEDPK ELAERFIKIF LEDMEALKVK PADIYPRVTD HIDDIIEFIG KLKEKGYAYE GSDGIYFEVK K FPEYGKLS ...String: MKHHHHHHGG SSGGLKVYNT LTKQKEEFKP LREGEVKMYV CGPTVYDYPH LGHARTYIAF DVIRRYLEHK GYTVLMVMNF TDIDDKIIK RARETGEDPK ELAERFIKIF LEDMEALKVK PADIYPRVTD HIDDIIEFIG KLKEKGYAYE GSDGIYFEVK K FPEYGKLS GVKIEDLQKG ARVEPGEGKK NPEDFALWKK AKPGEPKWDS PWGEGRPGWH IECSVMSSKY LGESFDIHGG GN DLIFPHH ENEIAQSEAC FGHEWVKYWL HTGFVMVKGE KMSKSLGNFV TIRELLKRYE PEVIRFFVLQ KHYRSPLEYT EEG LQHAKN NLQRLYNTLE NIRVALRNAE ISYTWGELEF KTYEIIREGK RKFYEAMDDD FNTAEALKAV FEVANAINKY LTEA NKPKE SILRKALEFF KIVSEVFGVF EDYFREETKE REESEKLIEL LVEVRKQLRK EKRYELADMI REELKKLGIQ LEDRG SETT WKRIIT UniProtKB: Cysteine--tRNA ligase |
-Macromolecule #2: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 2 / Number of copies: 2 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 283.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Temperature | Min: 186.0 K / Max: 192.0 K |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Average electron dose: 60.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
| Output model |  PDB-8qhp: |
 Movie
Movie Controller
Controller






 Z
Z Y
Y X
X