+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | S-layer protein SlaA from Sulfolobus acidocaldarius at pH 10.0 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  S-layer / dimer / S-layer / dimer /  N-glycosylation / N-glycosylation /  STRUCTURAL PROTEIN STRUCTURAL PROTEIN | |||||||||
| Function / homology |  S-layer / extracellular region / S-layer / extracellular region /  S-layer protein A S-layer protein A Function and homology information Function and homology information | |||||||||
| Biological species |    Sulfolobus acidocaldarius DSM 639 (acidophilic) Sulfolobus acidocaldarius DSM 639 (acidophilic) | |||||||||
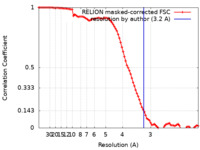
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.2 Å cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Gambelli L / Isupov MN / Daum B | |||||||||
| Funding support | European Union, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2024 Journal: Elife / Year: 2024Title: Structure of the two-component S-layer of the archaeon . Authors: Lavinia Gambelli / Mathew McLaren / Rebecca Conners / Kelly Sanders / Matthew C Gaines / Lewis Clark / Vicki A M Gold / Daniel Kattnig / Mateusz Sikora / Cyril Hanus / Michail N Isupov / Bertram Daum /     Abstract: Surface layers (S-layers) are resilient two-dimensional protein lattices that encapsulate many bacteria and most archaea. In archaea, S-layers usually form the only structural component of the cell ...Surface layers (S-layers) are resilient two-dimensional protein lattices that encapsulate many bacteria and most archaea. In archaea, S-layers usually form the only structural component of the cell wall and thus act as the final frontier between the cell and its environment. Therefore, S-layers are crucial for supporting microbial life. Notwithstanding their importance, little is known about archaeal S-layers at the atomic level. Here, we combined single-particle cryo electron microscopy, cryo electron tomography, and Alphafold2 predictions to generate an atomic model of the two-component S-layer of . The outer component of this S-layer (SlaA) is a flexible, highly glycosylated, and stable protein. Together with the inner and membrane-bound component (SlaB), they assemble into a porous and interwoven lattice. We hypothesise that jackknife-like conformational changes in SlaA play important roles in S-layer assembly. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15530.map.gz emd_15530.map.gz | 56.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15530-v30.xml emd-15530-v30.xml emd-15530.xml emd-15530.xml | 15.7 KB 15.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15530_fsc.xml emd_15530_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_15530.png emd_15530.png | 39.5 KB | ||
| Filedesc metadata |  emd-15530.cif.gz emd-15530.cif.gz | 6.8 KB | ||
| Others |  emd_15530_half_map_1.map.gz emd_15530_half_map_1.map.gz emd_15530_half_map_2.map.gz emd_15530_half_map_2.map.gz | 49.7 MB 49.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15530 http://ftp.pdbj.org/pub/emdb/structures/EMD-15530 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15530 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15530 | HTTPS FTP |
-Related structure data
| Related structure data |  8an2MC  7zcxC  8an3C  8qoxC  8qp0C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15530.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15530.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.054 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_15530_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_15530_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Monomeric S-layer protein SlaA with N-glycosylation
| Entire | Name: Monomeric S-layer protein SlaA with N-glycosylation |
|---|---|
| Components |
|
-Supramolecule #1: Monomeric S-layer protein SlaA with N-glycosylation
| Supramolecule | Name: Monomeric S-layer protein SlaA with N-glycosylation / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:    Sulfolobus acidocaldarius DSM 639 (acidophilic) Sulfolobus acidocaldarius DSM 639 (acidophilic) |
-Macromolecule #1: S-layer protein A
| Macromolecule | Name: S-layer protein A / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:    Sulfolobus acidocaldarius DSM 639 (acidophilic) Sulfolobus acidocaldarius DSM 639 (acidophilic)Strain: ATCC 33909 / DSM 639 / JCM 8929 / NBRC 15157 / NCIMB 11770 |
| Molecular weight | Theoretical: 151.078406 KDa |
| Recombinant expression | Organism:    Sulfolobus acidocaldarius DSM 639 (acidophilic) Sulfolobus acidocaldarius DSM 639 (acidophilic) |
| Sequence | String: MNKLVGLLVS SLFLASILIG IAPAITTTAL TPPVSAGGIQ AYLLTGSGAP ASGLVLFVVN VSNIQVSSSN VTNVISTVVS NIQINAKTE NAQTGATTGS VTVRFPTSGY NAYYDSVDKV VFVVVSFLYP YTTTSVNIPL SYLSKYLPGL LTAQPYDETG A QVTSVSST ...String: MNKLVGLLVS SLFLASILIG IAPAITTTAL TPPVSAGGIQ AYLLTGSGAP ASGLVLFVVN VSNIQVSSSN VTNVISTVVS NIQINAKTE NAQTGATTGS VTVRFPTSGY NAYYDSVDKV VFVVVSFLYP YTTTSVNIPL SYLSKYLPGL LTAQPYDETG A QVTSVSST PFGSLIDTST GQQILGTNPV LTSYNSYTTQ ANTNMQEGVV SGTLTSFTLG GQSFSGSTVP VILYAPFIFS NS PYQAGLY NPMQVNGNLG SLSSEAYYHP VIWGRALINT TLIDTYASGS VPFTFQLNYS VPGPLTINMA QLAWIASINN LPT SFTYLS YKFSNGYESF LGIISNSTQL TAGALTINPS GNFTINGKKF YVYLLVVGST NSTTPVEYVT KLVVEYPSST NFLP QGVTV TTSSNKYTLP VYEIGGPAGT TITLTGNWYS TPYTVQITVG STPTLTNYVS QILLKAVAYE GINVSTTQSP YYSTA ILST PPSEISITGS STITAQGKLT ATSASATVNL LTNATLTYEN IPLTQYSFNG IIVTPGYAAI NGTTAMAYVI GALYNK TSD YVLSFAGSQE PMQVMNNNLT EVTTLAPFGL TLLAPSVPAT ETGTSPLQLE FFTVPSTSYI ALVDFGLWGN LTSVTVS AY DTVNNKLSVN LGYFYGIVIP PSISTAPYNY QNFICPNNYV TVTIYDPDAV LDPYPSGSFT TSSLPLKYGN MNITGAVI F PGSSVYNPSG VFGYSNFNKG AAVTTFTYTA QSGPFSPVAL TGNTNYLSQY ADNNPTDNYY FIQTVNGMPV LMGGLSIVA SPVSASLPSS TSSPGFMYLL PSAAQVPSPL PGMATPNYNL NIYITYKIDG ATVGNNMING LYVASQNTLI YVVPNGSFVG SNIKLTYTT TDYAVLHYFY STGQYKVFKT VSVPNVTANL YFPSSTTPLY QLSVPLYLSE PYYGSPLPTY IGLGTNGTSL W NSPNYVLF GVSAVQQYLG FIKSISVTLS NGTTVVIPLT TSNMQTLFPQ LVGQELQACN GTFQFGISIT GLEKLLNLNV QQ LNNSILS VTYHDYVTGE TLTATTKLVA LSTLSLVAKG AGVVEFLLTA YPYTGNITFA PPWFIAENVV KQPFMTYSDL QFA KTNPSA ILSLSTVNIT VVGLGGKASV YYNSTSGQTV ITNIYGQTVA TLSGNVLPTL TELAAGNGTF TGSLQFTIVP NNTV VQIPS SLTKTSFAVY TNGSLAIVLN GKAYSLGPAG LFLLPFVTYT GSAIGANATA IITVSDGVGT STTQVPITAE NFTPI RLAP FQVPAQVPLP NAPKLKYEYN GSIVITPQQQ VLKIYVTSIL PYPQEFQIQA FVYEASQFNV HTGSPTAAPV YFSYSA VRA YPALGIGTSV PNLLVYVQLQ GISNLPAGKY VIVLSAVPFA GGPVLSEYPA QLIFTNVTLT Q UniProtKB:  S-layer protein A S-layer protein A |
-Macromolecule #8: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 8 / Number of copies: 3 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 10 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.4 µm / Nominal defocus min: 0.8 µm Bright-field microscopy / Nominal defocus max: 2.4 µm / Nominal defocus min: 0.8 µm |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 61.0 e/Å2 |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z
Z Y
Y X
X