[English] 日本語
 Yorodumi
Yorodumi- PDB-8fis: Structure of Bispecific CAP256V2LS-J3 Fab in complex with BG505 D... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8fis | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Bispecific CAP256V2LS-J3 Fab in complex with BG505 DS-SOSIP.664 | |||||||||
 Components Components |
| |||||||||
 Keywords Keywords |  VIRAL PROTEIN / CD4 / VIRAL PROTEIN / CD4 /  HIV-1 / SOSIP / HIV-1 / SOSIP /  Vaccine / Vaccine /  IMMUNE SYSTEM / IMMUNE SYSTEM /  llama / V2 apex / llama / V2 apex /  therapeutic / CAP256 / VRC26.25 therapeutic / CAP256 / VRC26.25 | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / positive regulation of establishment of T cell polarity / virus-mediated perturbation of host defense response / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell /  viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane /  viral envelope ...positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / positive regulation of establishment of T cell polarity / virus-mediated perturbation of host defense response / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral envelope ...positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / positive regulation of establishment of T cell polarity / virus-mediated perturbation of host defense response / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell /  viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane /  viral envelope / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity / identical protein binding / viral envelope / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity / identical protein binding /  plasma membrane plasma membraneSimilarity search - Function | |||||||||
| Biological species |    Human immunodeficiency virus 1 Human immunodeficiency virus 1  Lama glama (llama) Lama glama (llama)  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  ELECTRON MICROSCOPY / ELECTRON MICROSCOPY /  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.18 Å cryo EM / Resolution: 3.18 Å | |||||||||
 Authors Authors | Gorman, J. / Kwong, P.D. | |||||||||
| Funding support |  United States, 2items United States, 2items
| |||||||||
 Citation Citation |  Journal: MAbs / Year: 2023 Journal: MAbs / Year: 2023Title: Bispecific antibody CAP256.J3LS targets V2-apex and CD4-binding sites with high breadth and potency. Authors: Baoshan Zhang / Jason Gorman / Young D Kwon / Amarendra Pegu / Cara W Chao / Tracy Liu / Mangaiarkarasi Asokan / Michael F Bender / Tatsiana Bylund / Leland Damron / Deepika Gollapudi / ...Authors: Baoshan Zhang / Jason Gorman / Young D Kwon / Amarendra Pegu / Cara W Chao / Tracy Liu / Mangaiarkarasi Asokan / Michael F Bender / Tatsiana Bylund / Leland Damron / Deepika Gollapudi / Paula Lei / Yile Li / Cuiping Liu / Mark K Louder / Krisha McKee / Adam S Olia / Reda Rawi / Arne Schön / Shuishu Wang / Eun Sung Yang / Yongping Yang / Kevin Carlton / Nicole A Doria-Rose / Lawrence Shapiro / Michael S Seaman / John R Mascola / Peter D Kwong /  Abstract: Antibody CAP256-VRC26.25 targets the second hypervariable region (V2) at the apex of the HIV envelope (Env) trimer with extraordinary neutralization potency, although less than optimal breadth. To ...Antibody CAP256-VRC26.25 targets the second hypervariable region (V2) at the apex of the HIV envelope (Env) trimer with extraordinary neutralization potency, although less than optimal breadth. To improve breadth, we linked the light chain of CAP256V2LS, an optimized version of CAP256-VRC26.25 currently under clinical evaluation, to the llama nanobody J3, which has broad CD4-binding site-directed neutralization. The J3-linked bispecific antibody exhibited improved breadth and potency over both J3 and CAP256V2LS, indicative of synergistic neutralization. The cryo-EM structure of the bispecific antibody in complex with a prefusion-closed Env trimer revealed simultaneous binding of J3 and CAP256V2LS. We further optimized the pharmacokinetics of the bispecific antibody by reducing the net positive charge of J3. The optimized bispecific antibody, which we named CAP256.J3LS, had a half-life similar to CAP256V2LS in human FcRn knock-in mice and exhibited suitable auto-reactivity, manufacturability, and biophysical risk. CAP256.J3LS neutralized over 97% of a multiclade 208-strain panel (geometric mean concentration for 80% inhibition (IC) 0.079 μg/ml) and 100% of a 100-virus clade C panel (geometric mean IC of 0.05 μg/ml), suggesting its anti-HIV utility especially in regions where clade C dominates. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8fis.cif.gz 8fis.cif.gz | 511.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8fis.ent.gz pdb8fis.ent.gz | 425.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8fis.json.gz 8fis.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/fi/8fis https://data.pdbj.org/pub/pdb/validation_reports/fi/8fis ftp://data.pdbj.org/pub/pdb/validation_reports/fi/8fis ftp://data.pdbj.org/pub/pdb/validation_reports/fi/8fis | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 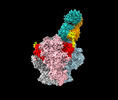 29209MC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
-Envelope glycoprotein ... , 2 types, 6 molecules ABECFG
| #1: Protein | Mass: 17146.482 Da / Num. of mol.: 3 / Fragment: UNP residues 509-661 Source method: isolated from a genetically manipulated source Source: (gene. exp.)    Human immunodeficiency virus 1 / Gene: env / Production host: Human immunodeficiency virus 1 / Gene: env / Production host:   Homo sapiens (human) / References: UniProt: Q2N0S6 Homo sapiens (human) / References: UniProt: Q2N0S6#2: Protein |  / Env polyprotein / Env polyproteinMass: 54086.324 Da / Num. of mol.: 3 / Fragment: UNP residues 30-510 Source method: isolated from a genetically manipulated source Source: (gene. exp.)    Human immunodeficiency virus 1 / Gene: env / Production host: Human immunodeficiency virus 1 / Gene: env / Production host:   Homo sapiens (human) / References: UniProt: Q2N0S6 Homo sapiens (human) / References: UniProt: Q2N0S6 |
|---|
-Protein , 2 types, 4 molecules JILH
| #3: Protein | Mass: 37355.211 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Lama glama (llama) / Production host: Lama glama (llama) / Production host:   Homo sapiens (human) Homo sapiens (human)#4: Protein | | Mass: 26863.039 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:   Homo sapiens (human) Homo sapiens (human) |
|---|
-Sugars , 6 types, 66 molecules 
| #5: Polysaccharide | beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta- ...beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose  / Mass: 586.542 Da / Num. of mol.: 8 / Mass: 586.542 Da / Num. of mol.: 8Source method: isolated from a genetically manipulated source #6: Polysaccharide | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose  / Mass: 424.401 Da / Num. of mol.: 22 / Mass: 424.401 Da / Num. of mol.: 22Source method: isolated from a genetically manipulated source #7: Polysaccharide |  / Mass: 748.682 Da / Num. of mol.: 3 / Mass: 748.682 Da / Num. of mol.: 3Source method: isolated from a genetically manipulated source #8: Polysaccharide |  / Mass: 748.682 Da / Num. of mol.: 3 / Mass: 748.682 Da / Num. of mol.: 3Source method: isolated from a genetically manipulated source #9: Polysaccharide | alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2- ...alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose  / Mass: 910.823 Da / Num. of mol.: 5 / Mass: 910.823 Da / Num. of mol.: 5Source method: isolated from a genetically manipulated source #10: Sugar | ChemComp-NAG /  N-Acetylglucosamine N-Acetylglucosamine |
|---|
-Details
| Has ligand of interest | N |
|---|
-Experimental details
-Experiment
| Experiment | Method:  ELECTRON MICROSCOPY ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method:  single particle reconstruction single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: CAP256V2LS-J3-3 Fab in complex with BG505 DS-SOSIP.664 Type: COMPLEX / Entity ID: #1-#4 / Source: RECOMBINANT |
|---|---|
| Molecular weight | Experimental value: NO |
| Source (natural) | Organism:    Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Source (recombinant) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Buffer solution | pH: 7.4 / Details: PBS |
| Specimen | Conc.: 2 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied : NO / Vitrification applied : NO / Vitrification applied : YES : YES |
| Specimen support | Grid type: C-flat-1.2/1.3 |
Vitrification | Instrument: FEI VITROBOT MARK IV / Cryogen name: ETHANE / Humidity: 90 % / Chamber temperature: 293 K |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source : :  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2500 nm / Nominal defocus min: 800 nm / C2 aperture diameter: 70 µm Bright-field microscopy / Nominal defocus max: 2500 nm / Nominal defocus min: 800 nm / C2 aperture diameter: 70 µm |
| Image recording | Average exposure time: 2 sec. / Electron dose: 51.19 e/Å2 / Detector mode: COUNTING / Film or detector model: GATAN K3 (6k x 4k) / Num. of grids imaged: 1 |
- Processing
Processing
| Software | Name: PHENIX / Version: 1.20_4459: / Classification: refinement | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EM software |
| ||||||||||||||||||||||||
CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||
| Symmetry | Point symmetry : C1 (asymmetric) : C1 (asymmetric) | ||||||||||||||||||||||||
3D reconstruction | Resolution: 3.18 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 118472 / Symmetry type: POINT | ||||||||||||||||||||||||
| Atomic model building | Protocol: FLEXIBLE FIT / Space: REAL | ||||||||||||||||||||||||
| Atomic model building | PDB-ID: 6NNF | ||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller


 PDBj
PDBj






