+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the human CCAN CENP-A alpha-satellite complex | ||||||||||||
 Map data Map data | Local resolution filtered cryosparc homogeneous refinement map | ||||||||||||
 Sample Sample |
| ||||||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of protein localization to kinetochore / Mis6-Sim4 complex / FANCM-MHF complex / centromere complex assembly / kinetochore organization / metaphase chromosome alignment / spindle attachment to meiosis I kinetochore / Fanconi anaemia nuclear complex / inner kinetochore /  kinetochore binding ...positive regulation of protein localization to kinetochore / Mis6-Sim4 complex / FANCM-MHF complex / centromere complex assembly / kinetochore organization / metaphase chromosome alignment / spindle attachment to meiosis I kinetochore / Fanconi anaemia nuclear complex / inner kinetochore / kinetochore binding ...positive regulation of protein localization to kinetochore / Mis6-Sim4 complex / FANCM-MHF complex / centromere complex assembly / kinetochore organization / metaphase chromosome alignment / spindle attachment to meiosis I kinetochore / Fanconi anaemia nuclear complex / inner kinetochore /  kinetochore binding / centromeric DNA binding / kinetochore binding / centromeric DNA binding /  sex differentiation / CENP-A containing chromatin assembly / resolution of meiotic recombination intermediates / chordate embryonic development / protein localization to chromosome, centromeric region / negative regulation of epithelial cell apoptotic process / sex differentiation / CENP-A containing chromatin assembly / resolution of meiotic recombination intermediates / chordate embryonic development / protein localization to chromosome, centromeric region / negative regulation of epithelial cell apoptotic process /  kinetochore assembly / attachment of mitotic spindle microtubules to kinetochore / condensed chromosome, centromeric region / replication fork processing / mitotic sister chromatid segregation / establishment of mitotic spindle orientation / mitotic cytokinesis / kinetochore assembly / attachment of mitotic spindle microtubules to kinetochore / condensed chromosome, centromeric region / replication fork processing / mitotic sister chromatid segregation / establishment of mitotic spindle orientation / mitotic cytokinesis /  chromosome, centromeric region / centriolar satellite / chromosome organization / negative regulation of megakaryocyte differentiation / protein localization to CENP-A containing chromatin / Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal / pericentric heterochromatin / interstrand cross-link repair / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / Packaging Of Telomere Ends / Mitotic Prometaphase / EML4 and NUDC in mitotic spindle formation / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Deposition of new CENPA-containing nucleosomes at the centromere / Resolution of Sister Chromatid Cohesion / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / Inhibition of DNA recombination at telomere / Meiotic synapsis / telomere organization / NRIF signals cell death from the nucleus / RNA Polymerase I Promoter Opening / Assembly of the ORC complex at the origin of replication / SUMOylation of chromatin organization proteins / mitotic spindle organization / chromosome, centromeric region / centriolar satellite / chromosome organization / negative regulation of megakaryocyte differentiation / protein localization to CENP-A containing chromatin / Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal / pericentric heterochromatin / interstrand cross-link repair / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / Packaging Of Telomere Ends / Mitotic Prometaphase / EML4 and NUDC in mitotic spindle formation / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Deposition of new CENPA-containing nucleosomes at the centromere / Resolution of Sister Chromatid Cohesion / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / Inhibition of DNA recombination at telomere / Meiotic synapsis / telomere organization / NRIF signals cell death from the nucleus / RNA Polymerase I Promoter Opening / Assembly of the ORC complex at the origin of replication / SUMOylation of chromatin organization proteins / mitotic spindle organization /  DNA methylation / Condensation of Prophase Chromosomes / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / Chromatin modifications during the maternal to zygotic transition (MZT) / SIRT1 negatively regulates rRNA expression / HCMV Late Events / DNA methylation / Condensation of Prophase Chromosomes / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / Chromatin modifications during the maternal to zygotic transition (MZT) / SIRT1 negatively regulates rRNA expression / HCMV Late Events /  innate immune response in mucosa / PRC2 methylates histones and DNA / positive regulation of protein ubiquitination / Defective pyroptosis / positive regulation of epithelial cell proliferation / innate immune response in mucosa / PRC2 methylates histones and DNA / positive regulation of protein ubiquitination / Defective pyroptosis / positive regulation of epithelial cell proliferation /  chromosome segregation / RHO GTPases Activate Formins / HDACs deacetylate histones / Fanconi Anemia Pathway / RNA Polymerase I Promoter Escape / Nonhomologous End-Joining (NHEJ) / Transcriptional regulation by small RNAs / Formation of the beta-catenin:TCF transactivating complex / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / PKR-mediated signaling / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / NoRC negatively regulates rRNA expression / B-WICH complex positively regulates rRNA expression / G2/M DNA damage checkpoint / HDMs demethylate histones / DNA Damage/Telomere Stress Induced Senescence / chromosome segregation / RHO GTPases Activate Formins / HDACs deacetylate histones / Fanconi Anemia Pathway / RNA Polymerase I Promoter Escape / Nonhomologous End-Joining (NHEJ) / Transcriptional regulation by small RNAs / Formation of the beta-catenin:TCF transactivating complex / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / PKR-mediated signaling / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / NoRC negatively regulates rRNA expression / B-WICH complex positively regulates rRNA expression / G2/M DNA damage checkpoint / HDMs demethylate histones / DNA Damage/Telomere Stress Induced Senescence /  kinetochore / Metalloprotease DUBs / PKMTs methylate histone lysines / RMTs methylate histone arginines / kinetochore / Metalloprotease DUBs / PKMTs methylate histone lysines / RMTs methylate histone arginines /  Meiotic recombination / Pre-NOTCH Transcription and Translation / Meiotic recombination / Pre-NOTCH Transcription and Translation /  nuclear matrix / nuclear matrix /  nucleosome assembly / Activation of anterior HOX genes in hindbrain development during early embryogenesis / HCMV Early Events / Transcriptional regulation of granulopoiesis / structural constituent of chromatin / Separation of Sister Chromatids / UCH proteinases / nucleosome assembly / Activation of anterior HOX genes in hindbrain development during early embryogenesis / HCMV Early Events / Transcriptional regulation of granulopoiesis / structural constituent of chromatin / Separation of Sister Chromatids / UCH proteinases /  nucleosome / antimicrobial humoral immune response mediated by antimicrobial peptide / nucleosome / antimicrobial humoral immune response mediated by antimicrobial peptide /  actin cytoskeleton / E3 ubiquitin ligases ubiquitinate target proteins / mitotic cell cycle / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / actin cytoskeleton / E3 ubiquitin ligases ubiquitinate target proteins / mitotic cell cycle / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks /  chromosome / RUNX1 regulates transcription of genes involved in differentiation of HSCs chromosome / RUNX1 regulates transcription of genes involved in differentiation of HSCsSimilarity search - Function | ||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||||||||
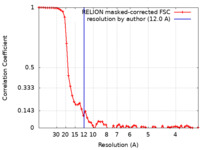
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 12.0 Å cryo EM / Resolution: 12.0 Å | ||||||||||||
 Authors Authors | Yatskevich S / Muir KW / Bellini D / Zhang Z / Yang J / Tischer T / Predin M / Dendooven T / McLaughlin SH / Barford D | ||||||||||||
| Funding support |  United Kingdom, United Kingdom,  Germany, 3 items Germany, 3 items
| ||||||||||||
 Citation Citation |  Journal: Science / Year: 2022 Journal: Science / Year: 2022Title: Structure of the human inner kinetochore bound to a centromeric CENP-A nucleosome. Authors: Stanislau Yatskevich / Kyle W Muir / Dom Bellini / Ziguo Zhang / Jing Yang / Thomas Tischer / Masa Predin / Tom Dendooven / Stephen H McLaughlin / David Barford /  Abstract: Kinetochores assemble onto specialized centromeric CENP-A (centromere protein A) nucleosomes (CENP-A) to mediate attachments between chromosomes and the mitotic spindle. We describe cryo-electron ...Kinetochores assemble onto specialized centromeric CENP-A (centromere protein A) nucleosomes (CENP-A) to mediate attachments between chromosomes and the mitotic spindle. We describe cryo-electron microscopy structures of the human inner kinetochore constitutive centromere associated network (CCAN) complex bound to CENP-A reconstituted onto α-satellite DNA. CCAN forms edge-on contacts with CENP-A, and a linker DNA segment of the α-satellite repeat emerges from the fully wrapped end of the nucleosome to thread through the central CENP-LN channel that tightly grips the DNA. The CENP-TWSX histone-fold module further augments DNA binding and partially wraps the linker DNA in a manner reminiscent of canonical nucleosomes. Our study suggests that the topological entrapment of the linker DNA by CCAN provides a robust mechanism by which kinetochores withstand both pushing and pulling forces exerted by the mitotic spindle. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14351.map.gz emd_14351.map.gz | 11.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14351-v30.xml emd-14351-v30.xml emd-14351.xml emd-14351.xml | 35.6 KB 35.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14351_fsc.xml emd_14351_fsc.xml | 6.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_14351.png emd_14351.png | 39.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14351 http://ftp.pdbj.org/pub/emdb/structures/EMD-14351 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14351 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14351 | HTTPS FTP |
-Related structure data
| Related structure data |  7ywxMC  7pb4C  7pb8C  7piiC  7pknC  7r5rC  7r5sC  7r5vC  7yyhC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14351.map.gz / Format: CCP4 / Size: 18.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14351.map.gz / Format: CCP4 / Size: 18.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local resolution filtered cryosparc homogeneous refinement map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.72 Å | ||||||||||||||||||||||||||||||||||||
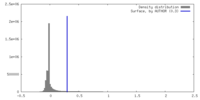
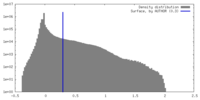
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
+Entire : CCAN-CENP-A inner centromere complex
+Supramolecule #1: CCAN-CENP-A inner centromere complex
+Macromolecule #1: Centromere protein H
+Macromolecule #2: Centromere protein I
+Macromolecule #3: Centromere protein K
+Macromolecule #4: Centromere protein L
+Macromolecule #5: Centromere protein M
+Macromolecule #6: Centromere protein N
+Macromolecule #7: Centromere protein O
+Macromolecule #8: Centromere protein Q
+Macromolecule #9: Centromere protein U
+Macromolecule #10: Centromere protein P
+Macromolecule #11: Centromere protein R
+Macromolecule #12: Centromere protein T
+Macromolecule #13: Centromere protein W
+Macromolecule #14: Centromere protein S
+Macromolecule #15: Centromere protein X
+Macromolecule #18: Histone H3-like centromeric protein A
+Macromolecule #19: Histone H4
+Macromolecule #20: Histone H2A type 1-C
+Macromolecule #21: Histone H2B type 1-C/E/F/G/I
+Macromolecule #22: Centromere protein C
+Macromolecule #16: DNA (171-MER)
+Macromolecule #17: DNA (171-MER)
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.6 µm / Nominal defocus min: 1.2 µm Bright-field microscopy / Nominal defocus max: 2.6 µm / Nominal defocus min: 1.2 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)