+Search query
-Structure paper
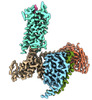
| Title | Structural insights into neurokinin 3 receptor activation by endogenous and analogue peptide agonists. |
|---|---|
| Journal, issue, pages | Cell Discov, Vol. 9, Issue 1, Page 66, Year 2023 |
| Publish date | Jun 30, 2023 |
 Authors Authors | Wenjing Sun / Fan Yang / Huanhuan Zhang / Qingning Yuan / Shenglong Ling / Yuanxia Wang / Pei Lv / Zelin Li / Yifan Luo / Dongsheng Liu / Wanchao Yin / Pan Shi / H Eric Xu / Changlin Tian /  |
| PubMed Abstract | Neurokinin 3 receptor (NK3R) is a tachykinin receptor essential for the hypothalamic-pituitary-gonadal axis. The endogenous peptide agonist neurokinin B (NKB) preferentially activates NK3R, while ...Neurokinin 3 receptor (NK3R) is a tachykinin receptor essential for the hypothalamic-pituitary-gonadal axis. The endogenous peptide agonist neurokinin B (NKB) preferentially activates NK3R, while substance P (SP) binds preferentially to NK1R. In addition, the SP analogue senktide more potently activates NK3R than NKB and SP. However, the mechanisms of preferential binding of peptide and NK3R activation remain elusive. Herein, we determined the cryogenic electron microscopy (cryo-EM) structures of the NK3R-G complex bound to NKB, SP and senktide. The three NK3R-G/peptide complexes utilize a class of noncanonical receptor activation mechanisms. Combining the structural analysis and functional assay illustrated that the consensus C-termini of the three peptide agonists share a conserved binding mode to NK3R, while the divergent N-termini of the peptides confer the preferential binding of the agonist to NK3R. In addition, the specific interactions between the N-terminus of senktide and the N-terminus and extracellular loops (ECL2 and ECL3) of NK3R lead to the improved activation displayed by senktide compared to SP and NKB. These findings pave the way to understand tachykinin receptor subtype selectivity and provide ideas to rationally develop drugs targeting NK3R. |
 External links External links |  Cell Discov / Cell Discov /  PubMed:37391393 / PubMed:37391393 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.8 - 3.0 Å |
| Structure data | EMDB-36144, PDB-8jbf: EMDB-36145, PDB-8jbg: EMDB-36146, PDB-8jbh: |
| Source |
|
 Keywords Keywords |  MEMBRANE PROTEIN / MEMBRANE PROTEIN /  G protein coupled receptor / G protein coupled receptor /  Neurokinin / Neurokinin /  Cryo-EM / Peptide Agonists Cryo-EM / Peptide Agonists |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers