+Search query
-Structure paper
| Title | McrD binds asymmetrically to methyl-coenzyme M reductase improving active-site accessibility during assembly. |
|---|---|
| Journal, issue, pages | Proc Natl Acad Sci U S A, Vol. 120, Issue 25, Page e2302815120, Year 2023 |
| Publish date | Jun 20, 2023 |
 Authors Authors | Grayson L Chadwick / Aaron M N Joiner / Sangeetha Ramesh / Douglas A Mitchell / Dipti D Nayak /  |
| PubMed Abstract | Methyl-coenzyme M reductase (MCR) catalyzes the formation of methane, and its activity accounts for nearly all biologically produced methane released into the atmosphere. The assembly of MCR is an ...Methyl-coenzyme M reductase (MCR) catalyzes the formation of methane, and its activity accounts for nearly all biologically produced methane released into the atmosphere. The assembly of MCR is an intricate process involving the installation of a complex set of posttranslational modifications and the unique Ni-containing tetrapyrrole called coenzyme F. Despite decades of research, details of MCR assembly remain largely unresolved. Here, we report the structural characterization of MCR in two intermediate states of assembly. These intermediate states lack one or both F cofactors and form complexes with the previously uncharacterized McrD protein. McrD is found to bind asymmetrically to MCR, displacing large regions of the alpha subunit and increasing active-site accessibility for the installation of F-shedding light on the assembly of MCR and the role of McrD therein. This work offers crucial information for the expression of MCR in a heterologous host and provides targets for the design of MCR inhibitors. |
 External links External links |  Proc Natl Acad Sci U S A / Proc Natl Acad Sci U S A /  PubMed:37307484 / PubMed:37307484 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 3.0 - 3.1 Å |
| Structure data | EMDB-29978, PDB-8gf5: EMDB-29979, PDB-8gf6: |
| Chemicals | 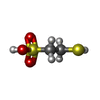 ChemComp-COM: 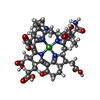 ChemComp-F43: 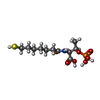 ChemComp-TP7: |
| Source |
|
 Keywords Keywords |  TRANSFERASE / TRANSFERASE /  methanogenesis / MCR complex methanogenesis / MCR complex |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers








