+Search query
-Structure paper
| Title | A wheat resistosome defines common principles of immune receptor channels. |
|---|---|
| Journal, issue, pages | Nature, Vol. 610, Issue 7932, Page 532-539, Year 2022 |
| Publish date | Sep 26, 2022 |
 Authors Authors | Alexander Förderer / Ertong Li / Aaron W Lawson / Ya-Nan Deng / Yue Sun / Elke Logemann / Xiaoxiao Zhang / Jie Wen / Zhifu Han / Junbiao Chang / Yuhang Chen / Paul Schulze-Lefert / Jijie Chai /   |
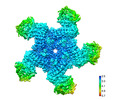
| PubMed Abstract | Plant intracellular nucleotide-binding leucine-rich repeat receptors (NLRs) detect pathogen effectors to trigger immune responses. Indirect recognition of a pathogen effector by the dicotyledonous ...Plant intracellular nucleotide-binding leucine-rich repeat receptors (NLRs) detect pathogen effectors to trigger immune responses. Indirect recognition of a pathogen effector by the dicotyledonous Arabidopsis thaliana coiled-coil domain containing NLR (CNL) ZAR1 induces the formation of a large hetero-oligomeric protein complex, termed the ZAR1 resistosome, which functions as a calcium channel required for ZAR1-mediated immunity. Whether the resistosome and channel activities are conserved among plant CNLs remains unknown. Here we report the cryo-electron microscopy structure of the wheat CNL Sr35 in complex with the effector AvrSr35 of the wheat stem rust pathogen. Direct effector binding to the leucine-rich repeats of Sr35 results in the formation of a pentameric Sr35-AvrSr35 complex, which we term the Sr35 resistosome. Wheat Sr35 and Arabidopsis ZAR1 resistosomes bear striking structural similarities, including an arginine cluster in the leucine-rich repeats domain not previously recognized as conserved, which co-occurs and forms intramolecular interactions with the 'EDVID' motif in the coiled-coil domain. Electrophysiological measurements show that the Sr35 resistosome exhibits non-selective cation channel activity. These structural insights allowed us to generate new variants of closely related wheat and barley orphan NLRs that recognize AvrSr35. Our data support the evolutionary conservation of CNL resistosomes in plants and demonstrate proof of principle for structure-based engineering of NLRs for crop improvement. |
 External links External links |  Nature / Nature /  PubMed:36163289 / PubMed:36163289 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 3.0 - 3.33 Å |
| Structure data |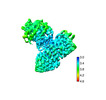  EMDB-33111: SR35-LRR with AvrSR35 EMDB-33112, PDB-7xc2: |
| Chemicals |  ChemComp-ATP: |
| Source |
|
 Keywords Keywords |  PLANT PROTEIN/INHIBITOR / LRR / CC / direct recognition / PLANT PROTEIN/INHIBITOR / LRR / CC / direct recognition /  PLANT PROTEIN-INHIBITOR COMPLEX PLANT PROTEIN-INHIBITOR COMPLEX |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers