-検索条件
-検索結果
検索 (著者・登録者: rosenthal & pb)の結果104件中、1から50件目までを表示しています

EMDB-18729: 
Cryo-EM structure of tetrameric human SAMHD1 with dApNHpp

EMDB-18730: 
Cryo-EM structure of tetrameric human SAMHD1 State I - Tense

EMDB-18731: 
Cryo-EM structure of tetrameric human SAMHD1 State II - Hemi-relaxed

EMDB-18732: 
Cryo-EM structure of tetrameric human SAMHD1 State III - Relaxed

EMDB-18733: 
Cryo-EM structure of tetrameric human SAMHD1 State IV - Depleted relaxed

EMDB-18734: 
Cryo-EM structure of tetrameric human SAMHD1 State V - Depleted relaxed

EMDB-17001: 
In-situ structure of the hexameric HEF trimers from influenza C viral particles
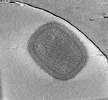
EMDB-18916: 
Cryotomogram of mature Vaccinia virus (WR) virion

EMDB-18917: 
Subtomogram average of the Vaccinia virus (WR) portal complex in mature virions

EMDB-18918: 
Subtomogram average of the Vaccinia virus (WR) A4/A10 palisade trimer in mature virions

EMDB-16150: 
8:1 binding of FcMR on IgM pentameric core

EMDB-16151: 
FcMR binding at subunit Fcu1 of IgM pentamer

EMDB-16152: 
FcMR binding at subunit Fcu3 of IgM pentamer

EMDB-15596: 
In situ subtomogram average of Vaccinia virus (WR) palisade, all virions

EMDB-15597: 
In situ subtomogram average of Vaccinia virus (WR) palisade, from CEVs

EMDB-15598: 
In situ subtomogram average of Vaccinia virus (WR) palisade, from IMVs

EMDB-15599: 
In situ subtomogram average of Vaccinia virus (WR) palisade, from IEVs

EMDB-15600: 
Cryotomogram of Vaccinia virus (WR) infected HeLa cell (immature virions)

EMDB-15601: 
Cryotomogram of Vaccinia virus (WR) infected HeLa cell (mature virions)

EMDB-15602: 
In situ subtomogram average of Vaccinia virus (WR) D13 lattice, on immature virions

EMDB-15181: 
Prefusion spike of SARS-CoV-2 (Wuhan), closed conformation
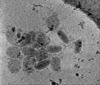
EMDB-15182: 
Electron cryotomography of SARS-CoV-2 virions

EMDB-15183: 
Whole SARS-CoV-2 (Wuhan) virion

EMDB-15185: 
Prefusion spike of SARS-CoV-2 (Wuhan), 1-RBD-up conformation

EMDB-13921: 
Cryo-EM structure of full-length human immunoglobulin M

EMDB-13922: 
Cryo-EM structure of human monomeric IgM-Fc

EMDB-15375: 
Cryo-EM structure of full-length human immunoglobulin M - F(ab')2 conformation 1

EMDB-15376: 
Cryo-EM structure of full-length human immunoglobulin M - F(ab')2 conformation 2

EMDB-15377: 
Cryo-EM structure of full-length human immunoglobulin M - F(ab')2 conformation 3

EMDB-15379: 
Cryo-EM structure of full-length human immunoglobulin M - F(ab')2 conformation 5

EMDB-15380: 
Cryo-EM structure of full-length human immunoglobulin M - F(ab')2 conformation 4

EMDB-14742: 
X-31 Hemagglutinin Precursor HA0 at pH 7.5

EMDB-14743: 
X-31 Hemagglutinin Precursor HA0 at pH 4.8

EMDB-14744: 
X-31 Hemagglutinin Precursor HA0 at pH 7.5 after reneutralization

EMDB-14225: 
Dissociated S1 domain of Alpha Variant SARS-CoV-2 Spike bound to ACE2 (Non-Uniform Refinement)

EMDB-14227: 
Dissociated S1 domain of Beta Variant SARS-CoV-2 Spike bound to ACE2 (Non-Uniform Refinement)

EMDB-14228: 
Dissociated S1 domain of Mink Variant SARS-CoV-2 Spike bound to ACE2 (Non-Uniform Refinement)

EMDB-14229: 
Alpha Variant SARS-CoV-2 Spike in Closed conformation

EMDB-14230: 
Alpha Variant SARS-CoV-2 Spike with 1 Erect RBD

EMDB-14232: 
Beta Variant SARS-CoV-2 Spike with 1 Erect RBD

EMDB-14233: 
Beta Variant SARS-CoV-2 Spike with 2 Erect RBDs

EMDB-14234: 
Mink Variant SARS-CoV-2 Spike in Closed conformation

EMDB-14235: 
Mink Variant SARS-CoV-2 Spike with 2 Erect RBDs

EMDB-14237: 
Mink Variant SARS-CoV-2 Spike with 1 Erect RBD

EMDB-14226: 
Dissociated S1 domain of Alpha Variant SARS-CoV-2 Spike bound to ACE2

EMDB-14231: 
Alpha Variant SARS-CoV-2 Spike with 2 Erect RBDs

EMDB-14236: 
Furin Cleaved Alpha Variant SARS-CoV-2 Spike in complex with 3 ACE2

EMDB-12585: 
Trimeric SARS-CoV-2 spike ectodomain in complex with biliverdin (closed conformation)

EMDB-12586: 
Trimeric SARS-CoV-2 spike ectodomain in complex with biliverdin (one RBD erect)

EMDB-12587: 
Trimeric SARS-CoV-2 spike ectodomain bound to P008_056 Fab
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
