-Search query
-Search result
Showing 1 - 50 of 60 items for (author: tu & lh)

EMDB-17819: 
XBB 1.0 RBD bound to P4J15 (Local)
Method: single particle / : Duhoo Y, Lau K
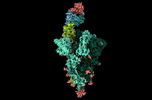
EMDB-17849: 
XBB 1.0 RBD bound to P4J15 (Global)
Method: single particle / : Duhoo Y, Lau K

EMDB-17850: 
SARS-CoV-2 XBB 1.0 closed conformation.
Method: single particle / : Duhoo Y, Lau K

EMDB-29530: 
SARS-CoV-2 XBB.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-29531: 
SARS-CoV-2 BQ.1.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-40240: 
SARS-CoV-2 BN.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-8fxb: 
SARS-CoV-2 XBB.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-8fxc: 
SARS-CoV-2 BQ.1.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-8s9g: 
SARS-CoV-2 BN.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-28558: 
SARS-CoV-2 BA.1 spike ectodomain trimer in complex with the S2X324 neutralizing antibody Fab fragment (local refinement of the RBD and S2X324)
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-28559: 
SARS-CoV-2 Omicron BA.1 spike ectodomain trimer in complex with the S2X324 neutralizing antibody Fab fragment
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-8erq: 
SARS-CoV-2 BA.1 spike ectodomain trimer in complex with the S2X324 neutralizing antibody Fab fragment (local refinement of the RBD and S2X324)
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-8err: 
SARS-CoV-2 Omicron BA.1 spike ectodomain trimer in complex with the S2X324 neutralizing antibody Fab fragment
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D
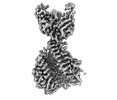
EMDB-27939: 
Structure of the human ACE2 receptor in complex with antibody Fab fragment, 05B04
Method: single particle / : Barnes CO

EMDB-27488: 
Cryo-EM structure of cystinosin in a cytosol-open state
Method: single particle / : Schmiege P, Li X

EMDB-27489: 
Cryo-EM structure of cystinosin in a lumen-open state
Method: single particle / : Schmiege P, Li X

EMDB-27490: 
Cryo-EM structure of cystine-bound cystinosin in a lumen-open state
Method: single particle / : Schmiege P, Li X

EMDB-27492: 
Cryo-EM structure of cystinosin N288K mutant in a cytosol-open state at pH5.0
Method: single particle / : Schmiege P, Li X

EMDB-27493: 
Cryo-EM structure of cystinosin N288K mutant in a cytosol-open state at pH7.5
Method: single particle / : Schmiege P, Li X

PDB-8dke: 
Cryo-EM structure of cystinosin in a cytosol-open state
Method: single particle / : Schmiege P, Li X

PDB-8dki: 
Cryo-EM structure of cystinosin in a lumen-open state
Method: single particle / : Schmiege P, Li X

PDB-8dkm: 
Cryo-EM structure of cystine-bound cystinosin in a lumen-open state
Method: single particle / : Schmiege P, Li X

PDB-8dkw: 
Cryo-EM structure of cystinosin N288K mutant in a cytosol-open state at pH5.0
Method: single particle / : Schmiege P, Li X

PDB-8dkx: 
Cryo-EM structure of cystinosin N288K mutant in a cytosol-open state at pH7.5
Method: single particle / : Schmiege P, Li X

EMDB-14777: 
Cryo-EM structure of archaic chaperone-usher Csu pilus of Acinetobacter baumannii
Method: helical / : Pakharukova N, Malmi H, Tuittila M, Paavilainen S, Ghosal D, Chang YW, Jensen GJ, Zavialov AV

PDB-7zl4: 
Cryo-EM structure of archaic chaperone-usher Csu pilus of Acinetobacter baumannii
Method: helical / : Pakharukova N, Malmi H, Tuittila M, Paavilainen S, Ghosal D, Chang YW, Jensen GJ, Zavialov AV
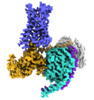
EMDB-24378: 
5-HT2AR bound to a novel agonist in complex with a mini-Gq protein and an active-state stabilizing single-chain variable fragment (scFv16) obtained by cryo-electron microscopy (cryoEM)
Method: single particle / : Barros-Alvarez X, Kim K, Panova O, Roth BL, Skiniotis G

PDB-7ran: 
5-HT2AR bound to a novel agonist in complex with a mini-Gq protein and an active-state stabilizing single-chain variable fragment (scFv16) obtained by cryo-electron microscopy (cryoEM)
Method: single particle / : Barros-Alvarez X, Kim K, Panova O, Roth BL, Skiniotis G

PDB-7ko8: 
Cryo-EM structure of the mature and infective Mayaro virus
Method: single particle / : Riberio-Filho HV, Coimbra LD, Cassago A, Rocha RPF, Padilha ACM, Schatz M, van Heel MG, Portugal RV, Trivella DBB, de Oliveira PSL, Marques RE

EMDB-22961: 
Cryo-EM structure of the mature and infective Mayaro virus
Method: single particle / : Ribeiro Filho H, Coimbra LD, Cassago A, Rocha RFP, Padilha AC, Trivella DBB, Schatz M, Oliveira PSL, Portugal RV, Marques RE, van Heel M

EMDB-12067: 
triplet microtubule from the proximal region of the pig sperm proximal centriole
Method: subtomogram averaging / : Leung MR, Zeev-Ben-Mordehai T

EMDB-12068: 
singlet microtubules from the pig sperm endpiece
Method: subtomogram averaging / : Leung MR, Zeev-Ben-Mordehai T

EMDB-12069: 
the central pair apparatus from pig sperm flagella
Method: subtomogram averaging / : Leung MR, Zeev-Ben-Mordehai T

EMDB-12070: 
the 96-nm axonemal repeat from pig sperm flagella (whole population)
Method: subtomogram averaging / : Leung MR, Zeev-Ben-Mordehai T

EMDB-12071: 
the 96-nm axonemal repeat from pig sperm flagella (+ODF class)
Method: subtomogram averaging / : Leung MR, Zeev-Ben-Mordehai T

EMDB-12072: 
the 96-nm axonemal repeat from pig sperm flagella (-ODF class)
Method: subtomogram averaging / : Leung MR, Zeev-Ben-Mordehai T

EMDB-12076: 
central pair apparatus from horse sperm flagella
Method: subtomogram averaging / : Leung MR, Zeev-Ben-Mordehai T

EMDB-12077: 
singlet microtubules from the horse sperm endpiece
Method: subtomogram averaging / : Leung MR, Zeev-Ben-Mordehai T

EMDB-12078: 
the 96-nm axonemal repeat from horse sperm flagella (whole population)
Method: subtomogram averaging / : Leung MR, Zeev-Ben-Mordehai T

EMDB-12079: 
the 96-nm axonemal repeat from horse sperm flagella (+ODF class)
Method: subtomogram averaging / : Leung MR, Zeev-Ben-Mordehai T

EMDB-12131: 
subtomogram average of the 96-nm axonemal repeat from horse sperm flagella (-ODF class)
Method: subtomogram averaging / : Leung MR, Zeev-Ben-Mordehai T

EMDB-12132: 
singlet microtubules from the mouse sperm endpiece
Method: subtomogram averaging / : Leung MR, Zeev-Ben-Mordehai T

EMDB-12133: 
the 96-nm axonemal repeat from mouse sperm flagella (whole population)
Method: subtomogram averaging / : Leung MR, Zeev-Ben-Mordehai T

EMDB-12134: 
the 96-nm axonemal repeat from mouse sperm flagella (+ODF class)
Method: subtomogram averaging / : Leung MR, Zeev-Ben-Mordehai T

EMDB-12135: 
the 96-nm axonemal repeat from mouse sperm flagella (-ODF class)
Method: subtomogram averaging / : Leung MR, Zeev-Ben-Mordehai T

EMDB-12136: 
the central pair apparatus from mouse sperm flagella
Method: subtomogram averaging / : Leung MR, Zeev-Ben-Mordehai T

EMDB-21097: 
Negative stain electron microscopy of PmPV2, a toxin member of the Membrane Attack Complex-Perforin (MACPF) from the eggs of Pomacea maculata
Method: single particle / : Otero LH, Brola TR, Giglio ML, Ituarte S, Dreon MS, Heras H

EMDB-10668: 
Structure of human ribosome in classical-PRE state
Method: single particle / : Bhaskar V, Schenk AD, Cavadini S, von Loeffelholz O, Natchiar SK, Klaholz BP, Chao JA
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search





 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
