-Search query
-Search result
Showing 1 - 50 of 151 items for (author: timothy & s & baker)

EMDB-41907: 
Computationally Designed, Expandable O4 Octahedral Handshake Nanocage
Method: single particle / : Weidle C, Borst A
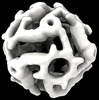
EMDB-42031: 
Computational Designed Nanocage O43_129_+8
Method: single particle / : Weidle C, Kibler RD

EMDB-43318: 
Twistless helix 12 repeat ring design R12B
Method: single particle / : Calise SJ, Kollman JM

EMDB-29974: 
Cryo-EM structure of synthetic tetrameric building block sC4
Method: single particle / : Redler RL, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-41364: 
CryoEM Structure of a Computationally Designed T3 Tetrahedral Nanocage
Method: single particle / : Weidle C, Borst AJ

EMDB-42906: 
Computational Designed Nanocage O43_129
Method: single particle / : Weidle C, Kibler RD

EMDB-42944: 
Computational Designed Nanocage O43_129_+4
Method: single particle / : Carr KD, Weidle C, Borst AJ

PDB-8gel: 
Cryo-EM structure of synthetic tetrameric building block sC4
Method: single particle / : Redler RL, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

PDB-8tl7: 
CryoEM Structure of a Computationally Designed T3 Tetrahedral Nanocage
Method: single particle / : Weidle C, Borst AJ

PDB-8v3b: 
Computational Designed Nanocage O43_129_+4
Method: single particle / : Carr KD, Weidle C, Borst AJ

EMDB-42601: 
CryoEM structure of Kappa Opioid Receptor bound to a semi-peptide and Gi1
Method: single particle / : Fay JF, Che T

EMDB-29026: 
CryoEM structure of Kappa Opioid Receptor bound to a semi-peptide and Gi1
Method: single particle / : Fay JF, Che T

PDB-8feg: 
CryoEM structure of Kappa Opioid Receptor bound to a semi-peptide and Gi1
Method: single particle / : Fay JF, Che T

EMDB-41153: 
Integrin alpha-v beta-8 in complex with minibinder B8_BP_dsulf
Method: single particle / : Campbell MG, Fernandez A, Roy A, Kraft J, Baker D

EMDB-41154: 
Integrin alpha-v beta-6 in complex with minibinder B6_BP_dslf
Method: single particle / : Campbell MG, Fernandez A, Roy A, Kraft J, Baker D

PDB-8tcf: 
Integrin alpha-v beta-8 in complex with minibinder B8_BP_dsulf
Method: single particle / : Campbell MG, Fernandez A, Roy A, Kraft J, Baker D

PDB-8tcg: 
Integrin alpha-v beta-6 in complex with minibinder B6_BP_dslf
Method: single particle / : Campbell MG, Fernandez A, Roy A, Kraft J, Baker D

EMDB-29915: 
CryoEM map of a de novo designed octahedral nanocage with programmable volume; design cage_O4_34
Method: single particle / : Kibler RD, Borst AJ

EMDB-40070: 
Cryo-EM map of synthetic cage_O3_10 reconstructed without symmetry (C1)
Method: single particle / : Coudray N, Redler R, Hsia Y, Huddy TF, Baker D, Ekiert D, Bhabha G

EMDB-40071: 
Cryo-EM map of synthetic cage_O3_10 reconstructed with O symmetry
Method: single particle / : Coudray N, Redler R, Hsia Y, Huddy TF, Baker D, Ekiert D, Bhabha G

EMDB-40073: 
Cryo-EM map of synthetic cage_T3_5 reconstructed without symmetry (C1), with 1 monomer missing (class 3.0)
Method: single particle / : Coudray N, Redler R, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-40074: 
Cryo-EM map of synthetic cage_T3_5 reconstructed with T symmetry
Method: single particle / : Coudray N, Redler R, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-40075: 
Cryo-EM map of synthetic cage_T3_5 reconstructed without symmetry (C1)
Method: single particle / : Coudray N, Redler R, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-40076: 
Cryo-EM map of synthetic cage_T3_5+2 reconstructed without symmetry (C1)
Method: single particle / : Coudray N, Redler R, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-40072: 
Cryo-EM map of synthetic cage_T3_5 reconstructed without symmetry (C1), with 1 trimer missing (class 3.1)
Method: single particle / : Coudray N, Redler R, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-23494: 
Cryo-EM of the SLFN12-PDE3A complex: PDE3A body refinement
Method: single particle / : Fuller JR, Garvie CW, Lemke CT

EMDB-23495: 
Cryo-EM of the SLFN12-PDE3A complex: Consensus subset model
Method: single particle / : Fuller JR, Garvie CW, Lemke CT

EMDB-23496: 
Cryo-EM of the SLFN12-PDE3A complex: SLFN12 body refinement
Method: single particle / : Fuller JR, Garvie CW, Lemke CT

PDB-7lrc: 
Cryo-EM of the SLFN12-PDE3A complex: PDE3A body refinement
Method: single particle / : Fuller JR, Garvie CW, Lemke CT

PDB-7lrd: 
Cryo-EM of the SLFN12-PDE3A complex: Consensus subset model
Method: single particle / : Fuller JR, Garvie CW, Lemke CT

PDB-7lre: 
Cryo-EM of the SLFN12-PDE3A complex: SLFN12 body refinement
Method: single particle / : Fuller JR, Garvie CW, Lemke CT

EMDB-20212: 
Cryo-EM structure of Bimetallic dodecameric cage design 3 (BMC3) from cytochrome cb562
Method: single particle / : Golub E, Subramanian RH, Yan X, Alberstein RG, Tezcan FA

PDB-6ovh: 
Cryo-EM structure of Bimetallic dodecameric cage design 3 (BMC3) from cytochrome cb562
Method: single particle / : Golub E, Subramanian RH, Yan X, Alberstein RG, Tezcan FA

EMDB-9105: 
Structure of a group II intron retroelement prior to DNA integration
Method: single particle / : Haack D, Yan X

EMDB-9106: 
Structure of a group II intron retroelement after DNA integration
Method: single particle / : Haack D, Yan X

PDB-6me0: 
Structure of a group II intron retroelement prior to DNA integration
Method: single particle / : Haack D, Yan X, Zhang C, Hingey J, Lyumkis D, Baker TS, Toor N

PDB-6mec: 
Structure of a group II intron retroelement after DNA integration
Method: single particle / : Haack D, Yan X, Zhang C, Hingey J, Lyumkis D, Baker TS, Toor N

EMDB-9600: 
The structure of CVA10 virus mature virion
Method: single particle / : Cui YX, Zheng QB, Zhu R, Xu LF, Li SW, Yan XD, Zhou ZH, Cheng T

EMDB-9601: 
The structure of CVA10 virus procapsid particle
Method: single particle / : Zhu R, Xu LF, Zheng QB, Cui YX, Li SW, Yan XD, Zhou ZH, Cheng T

EMDB-9602: 
The structure of CVA10 virus A-particle
Method: single particle / : Cui YX, Zheng QB, Zhu R, Xu LF, Li SW, Yan XD, Zhou ZH, Cheng T

EMDB-9603: 
The structure of CVA10 virus A-particle from its complex with Fab 2G8
Method: single particle / : Zhu R, Zheng QB, Xu LF, Cui YX, Li SW, Yan XD, Zhou ZH, Cheng T

EMDB-9604: 
The structure of CVA10 mature virion in complex with Fab 2G8
Method: single particle / : Zhu R, Zheng QB, Xu LF, Cui YX, Li SW, Yan XD, Zhou ZH, Cheng T

EMDB-9605: 
The structure of CVA10 procapsid from its complex with Fab 2G8
Method: single particle / : Zhu R, Zheng QB, Xu LF, Cui YX, Li SW, Yan XD, Zhou ZH, Cheng T

EMDB-9606: 
The structure of CVA10 virus A-particle
Method: single particle / : Zhu R, Zheng QB, Xu LF, Cui YX, Li SW, Yan XD, Zhou ZH, Cheng T

PDB-6acu: 
The structure of CVA10 virus mature virion
Method: single particle / : Cui YX, Zheng QB, Zhu R, Xu LF, Li SW, Yan XD, Zhou ZH, Cheng T

PDB-6acw: 
The structure of CVA10 virus procapsid particle
Method: single particle / : Zhu R, Xu LF, Zheng QB, Cui YX, Li SW, Yan XD, Zhou ZH, Cheng T

PDB-6acy: 
The structure of CVA10 virus A-particle
Method: single particle / : Cui YX, Zheng QB, Zhu R, Xu LF, Li SW, Yan XD, Zhou ZH, Cheng T

PDB-6acz: 
The structure of CVA10 virus A-particle from its complex with Fab 2G8
Method: single particle / : Zhu R, Zheng QB, Xu LF, Cui YX, Li SW, Yan XD, Zhou ZH, Cheng T

PDB-6ad0: 
The structure of CVA10 mature virion in complex with Fab 2G8
Method: single particle / : Zhu R, Zheng QB, Xu LF, Cui YX, Li SW, Yan XD, Zhou ZH, Cheng T
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
