-Search query
-Search result
Showing 1 - 50 of 722 items for (author: peng & ge)

EMDB-41363: 
Cryo-EM structure of DDB1deltaB-DDA1-DCAF5
Method: single particle / : Yue H, Hunkeler M, Roy Burman SS, Fischer ES

PDB-8tl6: 
Cryo-EM structure of DDB1deltaB-DDA1-DCAF5
Method: single particle / : Yue H, Hunkeler M, Roy Burman SS, Fischer ES

EMDB-17630: 
ABCB1 L335C mutant (mABCB1) in the inward facing state bound to AAC
Method: single particle / : Parey K, Januliene D, Gewering T, Moeller A, Vecchis D, Striednig B, Hilbi H, Schaefer LV, Kuprov I, Bordignon E, Seeger MA

PDB-8pee: 
ABCB1 L335C mutant (mABCB1) in the inward facing state bound to AAC
Method: single particle / : Parey K, Januliene D, Gewering T, Moeller A

EMDB-33347: 
Cryo-EM structure of S glycoprotein encoded by the Covid-19 mRNA vaccine candidate RQ3013 (Postfusion state)
Method: single particle / : Wu Z, Yu Z, Tan S, Lu J, Lu G, Lin J

PDB-7xog: 
Cryo-EM structure of S glycoprotein encoded by the Covid-19 mRNA vaccine candidate RQ3013 (Postfusion state)
Method: single particle / : Wu Z, Yu Z, Tan S, Lu J, Lu G, Lin J

EMDB-34314: 
SARS-CoV-2 RNA E-RTC complex with RMP-nsp9 and GMPPNP
Method: single particle / : Yan LM, Huang YC, Ge J, Liu ZY, Gao Y, Rao ZH, Lou ZY

EMDB-34316: 
SARS-CoV-2 E-RTC bound with MMP-nsp9 and GMPPNP
Method: single particle / : Yan LM, Rao ZH, Lou ZY

PDB-8gwk: 
SARS-CoV-2 RNA E-RTC complex with RMP-nsp9 and GMPPNP
Method: single particle / : Yan LM, Huang YC, Ge J, Liu ZY, Gao Y, Rao ZH, Lou ZY

PDB-8gwm: 
SARS-CoV-2 E-RTC bound with MMP-nsp9 and GMPPNP
Method: single particle / : Yan LM, Rao ZH, Lou ZY

EMDB-33346: 
Cryo-EM structure of S glycoprotein encoded by the Covid-19 mRNA vaccine candidate RQ3013 (Prefusion state)
Method: single particle / : Wu Z, Yu Z, Tan S, Lu J, Lu G, Lin J

PDB-7xoe: 
Cryo-EM structure of S glycoprotein encoded by the Covid-19 mRNA vaccine candidate RQ3013 (Prefusion state)
Method: single particle / : Wu Z, Yu Z, Tan S, Lu J, Lu G, Lin J

EMDB-38214: 
The structure of ASFV A137R
Method: single particle / : Li C, Song H, Gao GF
EMDB-37006: 
Cryo-EM structure of SARS-CoV-2 Delta RBD in complex with golden hamster ACE2 (local refinement)
Method: single particle / : Niu S, Zhao ZN, Chai Y, Gao GF

EMDB-37090: 
Cryo-EM structure of SARS-CoV-2 BA.3 RBD in complex with golden hamster ACE2 (local refinement)
Method: single particle / : Niu S, Zhao ZN, Chai Y, Gao GF

PDB-8ka8: 
Cryo-EM structure of SARS-CoV-2 Delta RBD in complex with golden hamster ACE2 (local refinement)
Method: single particle / : Niu S, Zhao ZN, Chai Y, Gao GF

PDB-8kc2: 
Cryo-EM structure of SARS-CoV-2 BA.3 RBD in complex with golden hamster ACE2 (local refinement)
Method: single particle / : Niu S, Zhao ZN, Chai Y, Gao GF
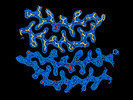

EMDB-41195: 
Tropomyosin-receptor kinase fused gene protein (TRK-fused gene protein; TFG) Low Complexity Domain (residues 237-327) G269V mutant, amyloid fiber
Method: helical / : Rosenberg GM, Sawaya MR, Boyer DR, Ge P, Abskharon R, Eisenberg DS
EMDB-41198: 
Tropomyosin-receptor kinase fused gene protein (TRK-fused gene protein; TFG) Low Complexity Domain (residues 237-327) P285L mutant, amyloid fiber
Method: helical / : Rosenberg GM, Sawaya MR, Boyer DR, Ge P, Abskharon R, Eisenberg DS

PDB-8teq: 
Tropomyosin-receptor kinase fused gene protein (TRK-fused gene protein; TFG) Low Complexity Domain (residues 237-327) G269V mutant, amyloid fiber
Method: helical / : Rosenberg GM, Sawaya MR, Boyer DR, Ge P, Abskharon R, Eisenberg DS

PDB-8ter: 
Tropomyosin-receptor kinase fused gene protein (TRK-fused gene protein; TFG) Low Complexity Domain (residues 237-327) P285L mutant, amyloid fiber
Method: helical / : Rosenberg GM, Sawaya MR, Boyer DR, Ge P, Abskharon R, Eisenberg DS

EMDB-37701: 
Cryo-EM structure of SARS-CoV-2 prototype RBD in complex with rabbit ACE2 (local refinement)
Method: single particle / : Li LJ, Shi KY, Yu GH, Gao GF

EMDB-37702: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.4/5 RBD in complex with rabbit ACE2 (local refinement)
Method: single particle / : Li LJ, Shi KY, Yu GH, Gao GF

EMDB-37703: 
Cryo-EM structure of SARS-CoV RBD in complex with rabbit ACE2
Method: single particle / : Li LJ, Shi KY, Yu GH, Gao GF

EMDB-37704: 
Cryo-EM map of SARS-CoV-2 prototype spike protein in complex with rabbit ACE2
Method: single particle / : Shi KY, Li LJ, Yu GH, Gao GF

EMDB-37706: 
Cryo-EM map of SARS-CoV-2 Omicron BA.4/5 spike protein in complex with rabbit ACE2
Method: single particle / : Li LJ, Shi KY, Yu GH, Gao GF

EMDB-38137: 
Cryo-EM map of SARS-CoV spike protein(6P) in complex with rabbit ACE2, 1-up state
Method: single particle / : Li LJ, Shi KY, Yu GH, Gao GF

EMDB-38144: 
Cryo-EM map of SARS-CoV spike protein(6P) in complex with rabbit ACE2, 2 RBD-up,1 ACE2-binding
Method: single particle / : Li LJ, Shi KY, Yu GH, Gao GF

EMDB-38152: 
Cryo-EM map of SARS-CoV spike protein(6P) in complex with rabbit ACE2, 3 RBD-up state
Method: single particle / : Li LJ, Shi KY, Yu GH, Gao GF

PDB-8wox: 
Cryo-EM structure of SARS-CoV-2 prototype RBD in complex with rabbit ACE2 (local refinement)
Method: single particle / : Li LJ, Shi KY, Yu GH, Gao GF

PDB-8woy: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.4/5 RBD in complex with rabbit ACE2 (local refinement)
Method: single particle / : Li LJ, Shi KY, Yu GH, Gao GF

PDB-8woz: 
Cryo-EM structure of SARS-CoV RBD in complex with rabbit ACE2
Method: single particle / : Li LJ, Shi KY, Yu GH, Gao GF

EMDB-35063: 
The complex structure of Omicron BA.4 RBD with BD604, S309, and S304
Method: single particle / : He QW, Xu ZP, Xie YF

EMDB-35064: 
SARS-CoV-2 Omicron BA.2 RBD complexed with BD-604 and S304 Fab
Method: single particle / : He QW, Xie Y

PDB-8hws: 
The complex structure of Omicron BA.4 RBD with BD604, S309, and S304
Method: single particle / : He QW, Xu ZP, Xie YF

PDB-8hwt: 
SARS-CoV-2 Omicron BA.2 RBD complexed with BD-604 and S304 Fab
Method: single particle / : He QW, Xie Y

EMDB-40450: 
Cryo-EM structure of CMKLR1 signaling complex
Method: single particle / : Zhang X, Zhang C

EMDB-34302: 
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Method: single particle / : Yan L, Rao Z, Lou Z

PDB-8gw1: 
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Method: single particle / : Yan L, Rao Z, Lou Z

EMDB-36622: 
The structure of EBOV L-VP35-RNA complex
Method: single particle / : Qi P, Yi S

EMDB-36623: 
The structure of EBOV L-VP35-RNA complex (conformation 1)
Method: single particle / : Qi P, Yi S

EMDB-36624: 
The structure of EBOV L-VP35-RNA complex (conformation 2)
Method: single particle / : Qi P, Yi S

PDB-8jsm: 
The structure of EBOV L-VP35-RNA complex (conformation 1)
Method: single particle / : Qi P, Yi S

PDB-8jsn: 
The structure of EBOV L-VP35-RNA complex (conformation 2)
Method: single particle / : Qi P, Yi S

EMDB-15687: 
The ABCB1 L335C mutant (mABCB1) in the Apo state
Method: single particle / : Parey K, Januliene D, Gewering T, Moeller A

PDB-8avy: 
The ABCB1 L335C mutant (mABCB1) in the Apo state
Method: single particle / : Parey K, Januliene D, Gewering T, Moeller A

EMDB-28014: 
HnRNPA2 D290V LCD PM3
Method: helical / : Eisenberg DS, Lu J, Ge P, Boyer DR
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search






 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
