-Search query
-Search result
Showing 1 - 50 of 62 items for (author: o & donnell & j)
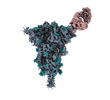
EMDB-40812: 
Structure of SARS-CoV-2 (HP-GSAS-Mut7) spike in complex with TXG-0078 Fab -Conformation 1
Method: single particle / : Bangaru S, Ward AB

EMDB-40813: 
Structure of SARS-CoV-2 (HP-GSAS-Mut7) spike in complex with TXG-0078 Fab -Conformation 2
Method: single particle / : Bangaru S, Ward AB

EMDB-40814: 
Local refinement of SARS-CoV-2 (HP-GSAS-Mut7) spike NTD in complex with TXG-0078 Fab
Method: single particle / : Bangaru B, Ward A

PDB-8swh: 
Local refinement of SARS-CoV-2 (HP-GSAS-Mut7) spike NTD in complex with TXG-0078 Fab
Method: single particle / : Bangaru B, Ward A

EMDB-42399: 
Cryo-EM structure of T4 Bacteriophage Clamp Loader with Sliding Clamp and DNA
Method: single particle / : Huang Y, Marcus K, Subramanian S, Gee LC, Gorday K, Ghaffari-Kashani S, Luo X, Zhang L, O'Donnell M, Kuriyan J

EMDB-42402: 
Cryo-EM structure of T4 Bacteriophage Clamp Loader with Sliding Clamp
Method: single particle / : Huang Y, Marcus K, Subramanian S, Gee LC, Gorday K, Ghaffari-Kashani S, Luo X, Zhang L, O'Donnell M, Kuriyan J

PDB-8unf: 
Cryo-EM structure of T4 Bacteriophage Clamp Loader with Sliding Clamp and DNA
Method: single particle / : Huang Y, Marcus K, Subramanian S, Gee LC, Gorday K, Ghaffari-Kashani S, Luo X, Zhang L, O'Donnell M, Subramanian S, Kuriyan J

PDB-8unh: 
Cryo-EM structure of T4 Bacteriophage Clamp Loader with Sliding Clamp
Method: single particle / : Huang Y, Marcus K, Subramanian S, Gee LC, Gorday K, Ghaffari-Kashani S, Luo X, Zhang L, O'Donnell M, Subramanian S, Kuriyan J

EMDB-26160: 
Structure of human serotonin transporter bound to small molecule '8090 in lipid nanodisc and NaCl
Method: single particle / : Singh I, Seth A

PDB-7txt: 
Structure of human serotonin transporter bound to small molecule '8090 in lipid nanodisc and NaCl
Method: single particle / : Singh I, Seth A, Billesboelle CB, Braz J, Rodriguiz RM, Roy K, Bekele B, Craik V, Huang XP, Boytsov D, Lak P, O'Donnell H, Sandtner W, Roth BL, Basbaum AI, Wetsel WC, Manglik A, Shoichet BK, Rudnick G

EMDB-27488: 
Cryo-EM structure of cystinosin in a cytosol-open state
Method: single particle / : Schmiege P, Li X

EMDB-27489: 
Cryo-EM structure of cystinosin in a lumen-open state
Method: single particle / : Schmiege P, Li X

EMDB-27490: 
Cryo-EM structure of cystine-bound cystinosin in a lumen-open state
Method: single particle / : Schmiege P, Li X

EMDB-27492: 
Cryo-EM structure of cystinosin N288K mutant in a cytosol-open state at pH5.0
Method: single particle / : Schmiege P, Li X

EMDB-27493: 
Cryo-EM structure of cystinosin N288K mutant in a cytosol-open state at pH7.5
Method: single particle / : Schmiege P, Li X

PDB-8dke: 
Cryo-EM structure of cystinosin in a cytosol-open state
Method: single particle / : Schmiege P, Li X

PDB-8dki: 
Cryo-EM structure of cystinosin in a lumen-open state
Method: single particle / : Schmiege P, Li X

PDB-8dkm: 
Cryo-EM structure of cystine-bound cystinosin in a lumen-open state
Method: single particle / : Schmiege P, Li X

PDB-8dkw: 
Cryo-EM structure of cystinosin N288K mutant in a cytosol-open state at pH5.0
Method: single particle / : Schmiege P, Li X

PDB-8dkx: 
Cryo-EM structure of cystinosin N288K mutant in a cytosol-open state at pH7.5
Method: single particle / : Schmiege P, Li X

EMDB-24310: 
The structure of human ABCG5-WT/ABCG8-I419E
Method: single particle / : Sun Y, Li X, Long T

EMDB-24311: 
The structure of human ABCG5-I529W/ABCG8-WT
Method: single particle / : Sun Y, Li X, Long T

EMDB-24312: 
The structure of human ABCG5/ABCG8 purified from yeast
Method: single particle / : Sun Y, Li X, Long T

EMDB-24313: 
The structure of human ABCG5/ABCG8 purified from mammalian cells
Method: single particle / : Sun Y, Li X, Long T

EMDB-24314: 
The structure of human ABCG5/ABCG8 supplemented with cholesterol
Method: single particle / : Sun Y, Li X, Long T

EMDB-24315: 
The structure of human ABCG1
Method: single particle / : Sun Y, Li X, Long T

EMDB-24316: 
The structure of human ABCG1 E242Q with cholesterol
Method: single particle / : Sun Y, Li X, Long T

EMDB-24317: 
The structure of human ABCG1 E242Q complexed with ATP
Method: single particle / : Sun Y, Li X, Long T

PDB-7r87: 
The structure of human ABCG5-WT/ABCG8-I419E
Method: single particle / : Sun Y, Li X, Long T

PDB-7r88: 
The structure of human ABCG5-I529W/ABCG8-WT
Method: single particle / : Sun Y, Li X, Long T

PDB-7r89: 
The structure of human ABCG5/ABCG8 purified from yeast
Method: single particle / : Sun Y, Li X, Long T

PDB-7r8a: 
The structure of human ABCG5/ABCG8 purified from mammalian cells
Method: single particle / : Sun Y, Li X, Long T

PDB-7r8b: 
The structure of human ABCG5/ABCG8 supplemented with cholesterol
Method: single particle / : Sun Y, Li X, Long T

PDB-7r8d: 
The structure of human ABCG1 E242Q with cholesterol
Method: single particle / : Sun Y, Li X, Long T

PDB-7r8e: 
The structure of human ABCG1 E242Q complexed with ATP
Method: single particle / : Sun Y, Li X, Long T

EMDB-12530: 
Subtomogram average of ChAdOx1 nCoV-19/AZD1222 derived SARS-CoV-2 spike glycoprotein
Method: subtomogram averaging / : Watanabe Y, Mendonca LM, Allen ER, Howe A, Lee M, Allen JD, Chawla H, Pulido D, Donnellan F, Davies H, Ulaszewska M, Belij-Rammerstorfer S, Morris S, Krebs AS, Dejnirattisai W, Mongkolsapaya J, Supasa P, Screaton GR, Green CM, Lambe T, Zhang P, Gilbert SC, Crispin M

PDB-6z3w: 
Human ER membrane protein complex
Method: single particle / : Hegde RS, O'Donnell JP

EMDB-10913: 
Structure of the phage STP1 host adhesion device (baseplate) by negative staining
Method: single particle / : Lavelle K, Goulet A, McDonnell B, Spinelli S, van Sinderen D, Mahony J, Cambillau C

EMDB-11058: 
Human ER Membrane protein Complex (EMC)
Method: single particle / : Phillips BP, O'Donnell JP

EMDB-8519: 
Structure of Eukaryotic CMG Helicase at a Replication Fork and Implications
Method: single particle / : Li B, Georgescu R, Yuan Z, Santos R, Sun J, Zhang D, Yurieva O, Li H, O'Donnell ME

PDB-5u8t: 
Structure of Eukaryotic CMG Helicase at a Replication Fork and Implications
Method: single particle / : Li B, Georgescu R, Yuan Z, Santos R, Sun J, Zhang D, Yurieva O, Li H, O'Donnell ME

EMDB-8518: 
Structure of eukaryotic CMG helicase at a replication fork
Method: single particle / : Li H, Li B, Georgescu R, Yuan Z, Santos R, Sun J, Zhang D, Yurieva O, O'Donnell ME

EMDB-8520: 
Structure of Eukaryotic CMG Helicase at a Replication Fork and Implications to Replisome Architecture and Origin Initiation
Method: single particle / : Georgescu R, Yuan Z, Bai L, Santos R, Sun J, Zhang D, Yurieva O, Li H, O'Donnell M

PDB-5u8s: 
Structure of eukaryotic CMG helicase at a replication fork
Method: single particle / : Li H, Li B, Georgescu R, Yuan Z, Santos R, Sun J, Zhang D, Yurieva O, O'Donnell ME

EMDB-6534: 
Structure of the eukaryotic replicative CMG helicase and pumpjack motion
Method: single particle / : Yuan Z, Bai L, Sun J, Georgescu RE, Liu J, O'Donnell ME, Li H

EMDB-6535: 
Structure of the eukaryotic replicative CMG helicase and pumpjack motion
Method: single particle / : Yuan Z, Bai L, Sun J, Georgescu RE, Liu J, O'Donnell ME, Li H

EMDB-6536: 
Structure of the eukaryotic replicative CMG helicase and pumpjack motion
Method: single particle / : Yuan Z, Bai L, Sun J, Georgescu RE, Liu J, O'Donnell ME, Li H

PDB-3jc5: 
Structure of the eukaryotic replicative CMG helicase and pumpjack motion
Method: single particle / : Li H, Bai L, Yuan Z, Sun J, Georgescu RE, Liu J, O'Donnell ME

PDB-3jc6: 
Structure of the eukaryotic replicative CMG helicase and pumpjack motion
Method: single particle / : Li H, Bai L, Yuan Z, Sun J, Georgescu RE, Liu J, O'Donnell ME
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
