-Search query
-Search result
Showing 1 - 50 of 85 items for (author: kaige & y)
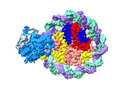
EMDB-37529: 
Structure of DDM1-nucleosome complex in the apo state
Method: single particle / : Liu Y, Zhang Z, Du J
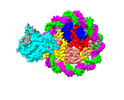
EMDB-37533: 
Structure of DDM1-nucleosome complex in ADP state
Method: single particle / : Liu Y, Zhang Z, Du J
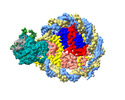
EMDB-37535: 
Structure of DDM1-nucleosome complex in ADP-BeFx state
Method: single particle / : Liu Y, Zhang Z, Du J
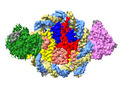

EMDB-37537: 
Structure of DDM1-nucleosome complex in the ADP-BeFx state with DDM1 bound to SHL2 and SHL-2
Method: single particle / : Liu Y, Zhang Z, Du J
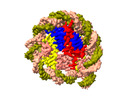
EMDB-37538: 
Structure of nucleosome core particle of Arabidopsis thaliana
Method: single particle / : Liu Y, Zhang Z, Du J

PDB-8wh5: 
Structure of DDM1-nucleosome complex in the apo state
Method: single particle / : Liu Y, Zhang Z, Du J

PDB-8wh8: 
Structure of DDM1-nucleosome complex in ADP state
Method: single particle / : Liu Y, Zhang Z, Du J

PDB-8wh9: 
Structure of DDM1-nucleosome complex in ADP-BeFx state
Method: single particle / : Liu Y, Zhang Z, Du J

PDB-8wha: 
Structure of DDM1-nucleosome complex in the ADP-BeFx state with DDM1 bound to SHL2 and SHL-2
Method: single particle / : Liu Y, Zhang Z, Du J

PDB-8whb: 
Structure of nucleosome core particle of Arabidopsis thaliana
Method: single particle / : Liu Y, Zhang Z, Du J

EMDB-37397: 
Cryo-EM structure of the ABCG25 E232Q mutant bound to ATP and Magnesium
Method: single particle / : Xin J, Yan KG

EMDB-37418: 
Cryo-EM structure of the ABCG25 bound to CHS
Method: single particle / : Xin J, Yan KG

EMDB-37426: 
Cryo-EM structure of the ABCG25 bound to ABA
Method: single particle / : Xin J, Yan KG

EMDB-37455: 
Cryo-EM structure of the ABCG25
Method: single particle / : Xin J, Yan KG

PDB-8wam: 
Cryo-EM structure of the ABCG25 E232Q mutant bound to ATP and Magnesium
Method: single particle / : Xin J, Yan KG

EMDB-34202: 
AK-42 inhibitor binding human ClC-2 TMD
Method: single particle / : Wang L
EMDB-35503: 
Cryo-EM structure of Stimulator of interferon genes
Method: single particle / : Lu DF, Shang GJ

EMDB-35504: 
Structure of Stimulator of interferon genes/ligand complex
Method: single particle / : Lu DF, Shang GJ

PDB-8ik0: 
Cryo-EM structure of Stimulator of interferon genes
Method: single particle / : Lu DF, Shang GJ

PDB-8ik3: 
Structure of Stimulator of interferon genes/ligand complex
Method: single particle / : Lu DF, Shang GJ

EMDB-34121: 
Cryo-EM structure of full-length Myosin Va in the autoinhibited state
Method: single particle / : Niu F, Wei Z

PDB-7yv9: 
Cryo-EM structure of full-length Myosin Va in the autoinhibited state
Method: single particle / : Niu F, Wei Z

EMDB-33832: 
CryoEM structure of Arabidopsis ROS1 in complex with TG mismatch dsDNA at 3.3 Angstroms resolution
Method: single particle / : Du X, Du J

EMDB-33835: 
CryoEM structure of Arabidopsis ROS1 in complex with 5mC-dsDNA at 3.1 Angstroms resolution
Method: single particle / : Du X, Du J

EMDB-33836: 
CryoEM structure of Arabidopsis ROS1 in complex with a covalent-linked reaction intermediate at 3.9 Angstroms resolution
Method: single particle / : Du X, Du J

PDB-7yho: 
CryoEM structure of Arabidopsis ROS1 in complex with TG mismatch dsDNA at 3.3 Angstroms resolution
Method: single particle / : Du X, Du J

PDB-7yhp: 
CryoEM structure of Arabidopsis ROS1 in complex with 5mC-dsDNA at 3.1 Angstroms resolution
Method: single particle / : Du X, Du J

PDB-7yhq: 
CryoEM structure of Arabidopsis ROS1 in complex with a covalent-linked reaction intermediate at 3.9 Angstroms resolution
Method: single particle / : Du X, Du J

EMDB-32944: 
The complex structure of Omicron BA.1 RBD with BD604, S309,and S304
Method: single particle / : Huang M, Xie YF

PDB-7x1m: 
The complex structure of Omicron BA.1 RBD with BD604, S309,and S304
Method: single particle / : Huang M, Xie YF, Qi JX

EMDB-10536: 
SUMOylated apoAPC/C with repositioned APC2 WHB domain
Method: single particle / : Barford D, Yatskevich S

EMDB-10538: 
SUMOylated apoAPC/C with density at APC2-APC4 interface
Method: single particle / : Barford D, Yatskevich S

PDB-6tnt: 
SUMOylated apoAPC/C with repositioned APC2 WHB domain
Method: single particle / : Barford D, Yatskevich S

EMDB-11626: 
Structure of inner kinetochore CCAN-Cenp-A complex (Cenp-HIKTW)
Method: single particle / : Yan K, Yang J, Zhang Z, Barford D

EMDB-30212: 
State 3 of pre50SH ribosome from rrmj knock out E.coli strain
Method: single particle / : Wang W, Li WQ, Ge XL, Yan KG, Mandava CS, Sanyal S, Gao N

EMDB-30213: 
State 2 of pre50SH ribosome from RrmJ knock out E.coli stain
Method: single particle / : Wei W, Wanqiu L, Xueliang G, Kaige Y, Chandra Sekhar M, Suparna S, Ning G

EMDB-30214: 
State 1 of pre50SH ribosome from RrmJ knock out E.coli stain
Method: single particle / : Wang W, Li WQ, Ge XL, Yan KG, Mandava CS, Sanyal S, Gao N

EMDB-30215: 
Mature 50S ribosomal subunit from RrmJ knock out E.coli strain
Method: single particle / : Wang W, Li WQ

PDB-7bv8: 
Mature 50S ribosomal subunit from RrmJ knock out E.coli strain
Method: single particle / : Wang W, Li WQ, Ge XL, Yan KG, Mandava CS, Sanyal S, Gao N

EMDB-4971: 
Structure of inner kinetochore CCAN complex
Method: single particle / : Yan K, Yang J, Zhang Z, McLaughlin SH, Chang L, Fasci D, Heck AJR, Barford D

EMDB-4579: 
Structure of inner kinetochore CCAN-Cenp-A complex
Method: single particle / : Yan K, Yang J

EMDB-4581: 
Structure of inner kinetochore CCAN complex with mask1
Method: single particle / : Yan K, Yang J

PDB-6qld: 
Structure of inner kinetochore CCAN-Cenp-A complex
Method: single particle / : Yan K, Yang J, Zhang Z, McLaughlin SH, Chang L, Fasci D, Heck AJR, Barford D

PDB-6qle: 
Structure of inner kinetochore CCAN complex
Method: single particle / : Yan K, Yang J, Zhang Z, McLaughlin SH, Chang L, Fasci D, Heck AJR, Barford D

PDB-6qlf: 
Structure of inner kinetochore CCAN complex with mask1
Method: single particle / : Yan K, Yang J, Zhang Z, McLaughlin SH, Chang L, Fasci D, Heck AJR, Barford D
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search








 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
