-Search query
-Search result
Showing all 38 items for (author: cuellar & j)

EMDB-16466: 
The structural architecture of alpha-synuclein oligomer
Method: single particle / : Cuellar J, Santos J, Pallares I, Ventura S, Valpuesta JM

EMDB-16528: 
3D reconstruction of alpha-synuclein oligomer-PSMa3 complex
Method: single particle / : Cuellar J, Santos J, Pallares I, Ventura S, Valpuesta JM

EMDB-14727: 
3D reconstruction of the cylindrical assembly of DnaJA2 by imposing D5 symmetry
Method: single particle / : Cuellar J, Velasco-Carneros L, Santiago C, Martin-Benito J, Valpuesta JM, Muga A

EMDB-14729: 
3D reconstruction of the cylindrical assembly of DnaJA2 without symmetry imposition
Method: single particle / : Cuellar J, Velasco-Carneros L, Santiago C, Martin-Benito J, Valpuesta JM, Muga A

EMDB-14736: 
3D reconstruction of the cylindrical assembly of DnaJA2 delta G/F by imposing D5 symmetry
Method: single particle / : Cuellar J, Velasco-Carneros L, Santiago C, Martin-Benito J, Valpuesta J, Muga A

PDB-7zhs: 
3D reconstruction of the cylindrical assembly of DnaJA2 delta G/F by imposing D5 symmetry
Method: single particle / : Cuellar J, Velasco-Carneros L, Santiago C, Martin-Benito J, Valpuesta J, Muga A

EMDB-14706: 
3D reconstruction of the cylindrical assembly of DnaJA2 delta G/F without symmetry imposition
Method: single particle / : Cuellar J, Velasco-Carneros L, Santiago C, Martin-Benito J, Muga A, Valpuesta JM

EMDB-13588: 
Gelsolin-free CCT
Method: single particle / : Cuellar J, Vallin J, Svanstrom A, Maestro-Lopez M, Bueno-Carrasco MT, Ludlam WG, Willardson BM, Valpuesta JM, Grantham J

EMDB-13177: 
Ser40 phosphorylated tyrosine hydroxylase
Method: single particle / : Bueno-Carrasco MT, Cuellar J

EMDB-13747: 
CCT-gelsolin complex
Method: single particle / : Cuellar J, Vallin J, Svanstrom A, Maestro-Lopez M, Bueno-Carrasco MT, Ludlam WG, Willardson BM, Valpuesta JM, Grantham J

EMDB-13442: 
Partial structure of tyrosine hydroxylase lacking the first 35 residues in complex with dopamine.
Method: single particle / : Bueno-Carrasco MT, Cuellar J

PDB-7pim: 
Partial structure of tyrosine hydroxylase lacking the first 35 residues in complex with dopamine.
Method: single particle / : Bueno-Carrasco MT, Cuellar J, Santiago C, Valpuesta JM, Martinez A, Flydal MI

EMDB-11309: 
Partial structure of tyrosine hydroxylase in complex with dopamine showing the catalytic domain and an alpha-helix from the regulatory domain involved in dopamine binding.
Method: single particle / : Bueno-Carrasco MT, Cuellar J

PDB-6zn2: 
Partial structure of tyrosine hydroxylase in complex with dopamine showing the catalytic domain and an alpha-helix from the regulatory domain involved in dopamine binding.
Method: single particle / : Bueno-Carrasco MT, Cuellar J, Santiago C, Valpuesta JM, Martinez A, Flydal MI

EMDB-11624: 
Full-length structure of the substrate-free tyrosine hydroxylase (apo-TH).
Method: single particle / : Bueno-Carrasco MT, Cuellar J

PDB-7a2g: 
Full-length structure of the substrate-free tyrosine hydroxylase (apo-TH).
Method: single particle / : Bueno-Carrasco MT, Cuellar J, Santiago C, Flydal MI, Martinez A, Valpuesta JM

EMDB-11467: 
Atomic model of the EM-based structure of the full-length tyrosine hydroxylase in complex with dopamine (residues 40-497) in which the regulatory domain (residues 40-165) has been included only with the backbone atoms
Method: single particle / : Bueno-Carrasco MT, Cuellar J, Santiago C, Valpuesta JM, Martinez A, Flydal MI

EMDB-11587: 
Partial structure of the substrate-free tyrosine hydroxylase (apo-TH).
Method: single particle / : Bueno-Carrasco MT, Cuellar J

PDB-6zvp: 
Atomic model of the EM-based structure of the full-length tyrosine hydroxylase in complex with dopamine (residues 40-497) in which the regulatory domain (residues 40-165) has been included only with the backbone atoms
Method: single particle / : Bueno-Carrasco MT, Cuellar J, Santiago C, Valpuesta JM, Martinez A, Flydal MI

PDB-6zzu: 
Partial structure of the substrate-free tyrosine hydroxylase (apo-TH).
Method: single particle / : Bueno-Carrasco MT, Cuellar J, Santiago C, Valpuesta JM, Martinez A, Flydal MI

EMDB-12749: 
In situ cryo-electron tomogram of a pyrenoid inside a Chlamydomonas reinhardtii cell
Method: electron tomography / : Cuellar LK, Schaffer M, Strauss M, Martinez-Sanchez A, Plitzko JM, Foerster F, Engel BD

EMDB-11992: 
the molecular sociology at the HeLa cell nuclear periphery
Method: electron tomography / : Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Kuhn-Cuellar L, Foerster F, Hyman A, Plitzko J, Baumeister W

EMDB-4489: 
Human CCT:mLST8 complex
Method: single particle / : Cuellar J, Santiago C, Ludlam WG, Bueno-Carrasco MT, Valpuesta JM, Willardson BM

EMDB-4503: 
Human substrate-free CCT
Method: single particle / : Cuellar J, Santiago C, Ludlam WG, Bueno-Carrasco MT, Valpuesta JM, Willardson BM

PDB-6qb8: 
Human CCT:mLST8 complex
Method: single particle / : Cuellar J, Santiago C, Ludlam WG, Bueno-Carrasco MT, Valpuesta JM, Willardson BM

EMDB-3694: 
In situ subtomogram average of Rubisco within the Chlamydomonas pyrenoid
Method: subtomogram averaging / : Cuellar LK, Schaffer M, Strauss M, Martinez-Sanchez A, Plitzko JM, Foerster F, Engel BD

EMDB-3442: 
Cryo-EM reconstructions of clathrin D6 cages
Method: single particle / : Sousa R, Liao HS, Cuellar J, Valpuesta JM, Jin AJ, Lafer EM

EMDB-4035: 
Cryo-EM reconstruction of clathrin D6 cages + full length Hsc70
Method: single particle / : Sousa R, Liao HS, Cuellar J, Valpuesta JM, Jin AJ, Lafer EM

EMDB-4036: 
Cryo-EM reconstruction of clathrin D6 cages + Hsc70 Delta C
Method: single particle / : Sousa R, Liao HS, Cuellar J, Valpuesta JM, Jin AJ, Lafer EM

EMDB-3232: 
The architecture of the S. pombe CCR4-NOT complex
Method: single particle / : Ukleja M, Cuellar J, Siwaszek A, Kasprzak JM, Czarnocki-Cieciura M, Bujnicki J, Dziembowski A, Valpuesta JM

EMDB-8054: 
In situ sub-tomogram average of HeLa nuclear pore complex from a single cell
Method: subtomogram averaging / : Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Kuhn-Cuellar L, Foerster F, Hyman A, Plitzko J, Baumeister W

EMDB-8055: 
In situ sub-tomogram average of HeLa nuclear pore complex
Method: subtomogram averaging / : Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Kuhn-Cuellar L, Foerster F, Hyman A, Plitzko J, Baumeister W

EMDB-8056: 
In situ sub-tomogram average of HeLa ER-associated ribosomes
Method: subtomogram averaging / : Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Kuhn-Cuellar L, Foerster F, Hyman A, Plitzko J, Baumeister W

EMDB-8057: 
In situ sub-tomogram average of HeLa cytosolic ribosomes
Method: subtomogram averaging / : Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Kuhn-Cuellar L, Foerster F, Hyman A, Plitzko J, Baumeister W

EMDB-2332: 
Structural insights into the chaperone activity of Hsp40: DnaJ binds and remodels RepE
Method: single particle / : Cuellar J, Perales-Calvo J, Muga A, Valpuesta JM, Moro F

EMDB-2333: 
Structural insights into the chaperone activity of Hsp40: DnaJ binds and remodels RepE
Method: single particle / : Cuellar J, Perales-Calvo J, Muga A, Valpuesta JM, Moro F

EMDB-2334: 
Structural insights into the chaperone activity of Hsp40: DnaJ binds and remodels RepE
Method: single particle / : Cuellar J, Perales-Calvo J, Muga A, Valpuesta JM, Moro F
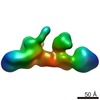
EMDB-2053: 
Electron Microscopy of THO complex
Method: single particle / : Pena A, Gewartowski K, Mroczek S, Cuellar J, Szykowska A, Prokop A, Czarnocki-Cieciura M, Piwowarski J, Tous C, Aguilera A, Carrascosa JL, Valpuesta JM, Dziembowski A
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
