[English] 日本語
 Yorodumi
Yorodumi- EMDB-5166: 3D reconstruction of a microtubule decorated with monomeric human... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5166 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
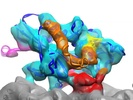
| Title | 3D reconstruction of a microtubule decorated with monomeric human kinesin (K349 construct) having AMPPNP bound in the nucleotide pocket. | |||||||||
 Map data Map data | 3D reconstruction of a microtubule decorated with monomeric human kinesin (K349 construct) with AMPPNP bound in the nucleotide pocket. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | kinesin / microtubule / atpase / molecular motor / myosin / mechanism / motility / cytoskeleton | |||||||||
| Biological species | unidentified (others) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 9.1 Å | |||||||||
 Authors Authors | Sindelar CV / Downing KH | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2010 Journal: Proc Natl Acad Sci U S A / Year: 2010Title: An atomic-level mechanism for activation of the kinesin molecular motors. Authors: Charles V Sindelar / Kenneth H Downing /  Abstract: Kinesin cytoskeletal motors convert the energy of ATP hydrolysis into stepping movement along microtubules. A partial model of this process has been derived from crystal structures, which show that ...Kinesin cytoskeletal motors convert the energy of ATP hydrolysis into stepping movement along microtubules. A partial model of this process has been derived from crystal structures, which show that movement of the motor domain relative to its major microtubule binding element, the switch II helix, is coupled to docking of kinesin's neck linker element along the motor domain. This docking would displace the cargo in the direction of travel and so contribute to a step. However, the crystal structures do not reveal how ATP binding and hydrolysis govern this series of events. We used cryoelectron microscopy to derive 8-9 A-resolution maps of four nucleotide states encompassing the microtubule-attached kinetic cycle of a kinesin motor. The exceptionally high quality of these maps allowed us to build in crystallographically determined conformations of kinesin's key subcomponents, yielding novel arrangements of kinesin's switch II helix and nucleotide-sensing switch loops. The resulting atomic models reveal a seesaw mechanism in which the switch loops, triggered by ATP binding, propel their side of the motor domain down and thereby elicit docking of the neck linker on the opposite side of the seesaw. Microtubules engage the seesaw mechanism by stabilizing the formation of extra turns at the N terminus of the switch II helix, which then serve as an anchor for the switch loops as they modulate the seesaw angle. These observations explain how microtubules activate kinesin's ATP-sensing machinery to promote cargo displacement and inform the mechanism of kinesin's ancestral relative, myosin. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5166.map.gz emd_5166.map.gz | 154.9 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5166-v30.xml emd-5166-v30.xml emd-5166.xml emd-5166.xml | 10 KB 10 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5166.jpg emd_5166.jpg | 78.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5166 http://ftp.pdbj.org/pub/emdb/structures/EMD-5166 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5166 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5166 | HTTPS FTP |
-Validation report
| Summary document |  emd_5166_validation.pdf.gz emd_5166_validation.pdf.gz | 78.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5166_full_validation.pdf.gz emd_5166_full_validation.pdf.gz | 77.7 KB | Display | |
| Data in XML |  emd_5166_validation.xml.gz emd_5166_validation.xml.gz | 493 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5166 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5166 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5166 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5166 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5166.map.gz / Format: CCP4 / Size: 165 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5166.map.gz / Format: CCP4 / Size: 165 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D reconstruction of a microtubule decorated with monomeric human kinesin (K349 construct) with AMPPNP bound in the nucleotide pocket. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.1167 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Microtubule decorated with monomeric human kinesin (K349 construc...
| Entire | Name: Microtubule decorated with monomeric human kinesin (K349 construct) having AMPPNP bound in the nucleotide pocket. |
|---|---|
| Components |
|
-Supramolecule #1000: Microtubule decorated with monomeric human kinesin (K349 construc...
| Supramolecule | Name: Microtubule decorated with monomeric human kinesin (K349 construct) having AMPPNP bound in the nucleotide pocket. type: sample / ID: 1000 / Number unique components: 2 |
|---|
-Macromolecule #1: K349
| Macromolecule | Name: K349 / type: protein_or_peptide / ID: 1 / Name.synonym: Monomeric human kinesin construct / Recombinant expression: Yes / Database: NCBI |
|---|---|
| Source (natural) | Organism: unidentified (others) |
-Macromolecule #2: Microtubule
| Macromolecule | Name: Microtubule / type: protein_or_peptide / ID: 2 / Name.synonym: Microtubule / Recombinant expression: Yes / Database: NCBI |
|---|---|
| Source (natural) | Organism: unidentified (others) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 6.8 / Details: 25mM Pipes, 25mM NaCl, 2mM MgCl2, 1mM EGTA |
|---|---|
| Grid | Details: 300 mesh copper grid with homemade holey carbon |
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER / Details: Vitrification instrument: Homemade |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 4000EX |
|---|---|
| Temperature | Average: 105 K |
| Date | Jan 21, 2008 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: NIKON SUPER COOLSCAN 9000 / Digitization - Sampling interval: 6.35 µm / Number real images: 98 / Average electron dose: 15 e/Å2 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 400 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 4.1 mm / Nominal defocus max: 1.6 µm / Nominal defocus min: 0.7 µm / Nominal magnification: 60000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 9.1 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: SPIDER,FREALIGN Details: Approximately 190,000 asymmetric units were averaged in the final reconstruction. |
|---|---|
| CTF correction | Details: done within FREALIGN |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name:  UCSF Chimera UCSF Chimera |
| Details | Protocol: Rigid Body. Regions in the crystal structure whose structure were found to differ from the EM map were excluded from the fitting procedure. These regions corresponded to the switch loops, helices alpha 3 and alpha 4 (switch II helix), and the neck linker. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: Cross correlation |
 Movie
Movie Controller
Controller