[English] 日本語
 Yorodumi
Yorodumi- PDB-4d2x: Negative-stain electron microscopy of E. coli ClpB of Y503D hyper... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4d2x | ||||||
|---|---|---|---|---|---|---|---|
| Title | Negative-stain electron microscopy of E. coli ClpB of Y503D hyperactive mutant (BAP form bound to ClpP) | ||||||
 Components Components | CHAPERONE PROTEIN CLPB | ||||||
 Keywords Keywords | CHAPERONE / DISAGGREGASE / CLPB / BAP / Y503D HYPERACTIVE MUTANT / COILED- COIL DOMAIN | ||||||
| Function / homology |  Function and homology information Function and homology informationcellular response to heat / protein refolding / response to heat / ATP hydrolysis activity / ATP binding / identical protein binding / membrane / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / negative staining / Resolution: 20 Å | ||||||
 Authors Authors | Carroni, M. / Kummer, E. / Oguchi, Y. / Clare, D.K. / Wendler, P. / Sinning, I. / Kopp, J. / Mogk, A. / Bukau, B. / Saibil, H.R. | ||||||
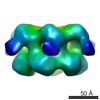
 Citation Citation |  Journal: Elife / Year: 2014 Journal: Elife / Year: 2014Title: Head-to-tail interactions of the coiled-coil domains regulate ClpB activity and cooperation with Hsp70 in protein disaggregation. Authors: Marta Carroni / Eva Kummer / Yuki Oguchi / Petra Wendler / Daniel K Clare / Irmgard Sinning / Jürgen Kopp / Axel Mogk / Bernd Bukau / Helen R Saibil /   Abstract: The hexameric AAA+ chaperone ClpB reactivates aggregated proteins in cooperation with the Hsp70 system. Essential for disaggregation, the ClpB middle domain (MD) is a coiled-coil propeller that binds ...The hexameric AAA+ chaperone ClpB reactivates aggregated proteins in cooperation with the Hsp70 system. Essential for disaggregation, the ClpB middle domain (MD) is a coiled-coil propeller that binds Hsp70. Although the ClpB subunit structure is known, positioning of the MD in the hexamer and its mechanism of action are unclear. We obtained electron microscopy (EM) structures of the BAP variant of ClpB that binds the protease ClpP, clearly revealing MD density on the surface of the ClpB ring. Mutant analysis and asymmetric reconstructions show that MDs adopt diverse positions in a single ClpB hexamer. Adjacent, horizontally oriented MDs form head-to-tail contacts and repress ClpB activity by preventing Hsp70 interaction. Tilting of the MD breaks this contact, allowing Hsp70 binding, and releasing the contact in adjacent subunits. Our data suggest a wavelike activation of ClpB subunits around the ring.DOI: http://dx.doi.org/10.7554/eLife.02481.001. | ||||||
| History |
| ||||||
| Remark 650 | HELIX DETERMINATION METHOD: AUTHOR PROVIDED. | ||||||
| Remark 700 | SHEET DETERMINATION METHOD: AUTHOR PROVIDED. |
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4d2x.cif.gz 4d2x.cif.gz | 492.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4d2x.ent.gz pdb4d2x.ent.gz | 285.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4d2x.json.gz 4d2x.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  4d2x_validation.pdf.gz 4d2x_validation.pdf.gz | 698.9 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  4d2x_full_validation.pdf.gz 4d2x_full_validation.pdf.gz | 708.3 KB | Display | |
| Data in XML |  4d2x_validation.xml.gz 4d2x_validation.xml.gz | 80.7 KB | Display | |
| Data in CIF |  4d2x_validation.cif.gz 4d2x_validation.cif.gz | 138.7 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/d2/4d2x https://data.pdbj.org/pub/pdb/validation_reports/d2/4d2x ftp://data.pdbj.org/pub/pdb/validation_reports/d2/4d2x ftp://data.pdbj.org/pub/pdb/validation_reports/d2/4d2x | HTTPS FTP |
-Related structure data
| Related structure data | 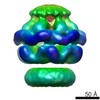 2559MC  2555C  2556C  2557C  2558C 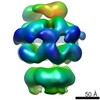 2560C  2561C  2562C  2563C  4ciuC  4d2qC  4d2uC C: citing same article ( M: map data used to model this data |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 96835.086 Da / Num. of mol.: 6 Source method: isolated from a genetically manipulated source Details: THE PROTEIN IS ENGINEERED TO BIND TO CLPP. / Source: (gene. exp.)   |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: BAP FORM OF CLPB (Y503D MUTANT) WITH ATPGS / Type: COMPLEX |
|---|---|
| Buffer solution | pH: 7.5 |
| Specimen | Conc.: 0.03 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied: YES / Vitrification applied: NO |
| EM staining | Type: NEGATIVE / Material: uranyl acetate |
| Specimen support | Details: CARBON |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TECNAI F20 / Date: Jul 20, 2011 |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal magnification: 50000 X / Calibrated magnification: 68000 X / Nominal defocus max: 1200 nm / Nominal defocus min: 500 nm / Cs: 2 mm |
| Image recording | Electron dose: 20 e/Å2 / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) |
| Image scans | Num. digital images: 108 |
- Processing
Processing
| EM software |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CTF correction | Details: PHASE FLIPPING ENTIRE FRAME | ||||||||||||
| Symmetry | Point symmetry: C6 (6 fold cyclic) | ||||||||||||
| 3D reconstruction | Method: ANGULAR RECONSTITUTION AND PROJECTION MATCHING / Resolution: 20 Å / Num. of particles: 9436 Magnification calibration: SUBMISSION BASED ON EXPERIMENTAL DATA FROM EMDB EMD-2559. (DEPOSITION ID: 12244). Symmetry type: POINT | ||||||||||||
| Atomic model building | Protocol: RIGID BODY FIT / Details: METHOD--RIGID BODY | ||||||||||||
| Refinement | Highest resolution: 20 Å | ||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 20 Å
|
 Movie
Movie Controller
Controller







 PDBj
PDBj



