| Entry | Database: PDB / ID: 2yh5
|
|---|
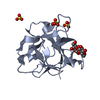
| Title | Structure of the C-terminal domain of BamC |
|---|
 Components Components | DAPX PROTEIN |
|---|
 Keywords Keywords | LIPID BINDING PROTEIN / LIPOPROTEIN / BAM COMPLEX |
|---|
| Function / homology |  Function and homology information Function and homology information
Bam protein complex / Gram-negative-bacterium-type cell outer membrane assembly / Secretion of toxins / protein insertion into membrane / cell surface / identical protein binding / membraneSimilarity search - Function Outer membrane protein assembly factor BamC / Outer membrane protein assembly factor BamC / NlpB/DapX lipoprotein / Outer membrane protein assembly factor BamC, C-terminal / NlpB/DapX lipoprotein / TATA-Binding Protein / Prokaryotic membrane lipoprotein lipid attachment site profile. / 2-Layer Sandwich / Alpha BetaSimilarity search - Domain/homology |
|---|
| Biological species |   ESCHERICHIA COLI (E. coli) ESCHERICHIA COLI (E. coli) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MIRAS / Resolution: 1.25 Å MIRAS / Resolution: 1.25 Å |
|---|
 Authors Authors | Zeth, K. / Albrecht, R. |
|---|
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2011 Journal: J.Biol.Chem. / Year: 2011
Title: Structural Basis of Outer Membrane Protein Biogenesis in Bacteria.
Authors: Albrecht, R. / Zeth, K. |
|---|
| History | | Deposition | Apr 27, 2011 | Deposition site: PDBE / Processing site: PDBE |
|---|
| Revision 1.0 | May 25, 2011 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | Aug 10, 2011 | Group: Database references / Version format compliance |
|---|
| Revision 1.2 | Feb 5, 2014 | Group: Refinement description |
|---|
| Revision 1.3 | May 8, 2024 | Group: Data collection / Database references ...Data collection / Database references / Derived calculations / Other
Category: chem_comp_atom / chem_comp_bond ...chem_comp_atom / chem_comp_bond / database_2 / pdbx_database_status / struct_site
Item: _database_2.pdbx_DOI / _database_2.pdbx_database_accession ..._database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_database_status.status_code_sf / _struct_site.pdbx_auth_asym_id / _struct_site.pdbx_auth_comp_id / _struct_site.pdbx_auth_seq_id |
|---|
|
|---|
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MIRAS / Resolution: 1.25 Å
MIRAS / Resolution: 1.25 Å  Authors
Authors Citation
Citation Journal: J.Biol.Chem. / Year: 2011
Journal: J.Biol.Chem. / Year: 2011 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 2yh5.cif.gz
2yh5.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb2yh5.ent.gz
pdb2yh5.ent.gz PDB format
PDB format 2yh5.json.gz
2yh5.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/yh/2yh5
https://data.pdbj.org/pub/pdb/validation_reports/yh/2yh5 ftp://data.pdbj.org/pub/pdb/validation_reports/yh/2yh5
ftp://data.pdbj.org/pub/pdb/validation_reports/yh/2yh5 Links
Links Assembly
Assembly
 Components
Components

 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  SLS
SLS  / Beamline: X10SA / Wavelength: 1
/ Beamline: X10SA / Wavelength: 1  Processing
Processing MIRAS
MIRAS Movie
Movie Controller
Controller
















 PDBj
PDBj






