 Yorodumi
Yorodumi+ Open data
Open data
- Basic information
Basic information
| Entry | 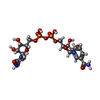 Database: PDB chemical components / ID: A7R Database: PDB chemical components / ID: A7R |
|---|---|
| Name | Name: [[(2S,3S,4R,5S)-5-(3-aminocarbonylpyridin-1-ium-1-yl)-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-oxidanyl-phosphoryl] [(2S,3S,4R,5S)-5-(4-azanyl-2-oxidanylidene-pyrimidin-1-yl)-3,4-bis(oxidanyl) ...Name: [[( |
-Chemical information
| Composition |  | ||||
|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAIN / Three letter code: A7R / Ideal coordinates details: Corina / Model coordinates PDB-ID: 6IH3 | ||||
| History |
| ||||
 External links External links |  UniChem / UniChem /  PubChem / PubChem /  ChemSpider / ChemSpider /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| CACTVS 3.385 | | OpenEye OEToolkits 2.0.6 | |
|---|
-SMILES CANONICAL
| CACTVS 3.385 | | OpenEye OEToolkits 2.0.6 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-PDB entries
Showing all 5 items

PDB-6ih3: 
Crystal structure of Phosphite Dehydrogenase from Ralstonia sp. 4506 in complex with non-natural cofactor Nicotinamide Cytosine Dinucleotide

PDB-6ih5: 
Crystal structure of Phosphite Dehydrogenase mutant I151R/P176E from Ralstonia sp. 4506 in complex with non-natural cofactor Nicotinamide Cytosine dinucleotide

PDB-6ih6: 
Phosphite Dehydrogenase mutant I151R/P176R/M207A from Ralstonia sp. 4506 in complex with non-natural cofactor Nicotinamide Cytosine dinucleotide

PDB-6juj: 
Crystal structure of Formate dehydrogenase mutant V198I/C256I/P260S/E261P/S381N/S383F from Pseudomonas sp. 101in complex with non-natural cofactor Nicotinamide Cytosine Dinucleotide

PDB-6juk: 
Crystal structure of Formate dehydrogenase mutant C256I/E261P/S381I from Pseudomonas sp. 101 in complex with non-natural cofactor Nicotinamide Cytosine Dinucleotide
 Movie
Movie Controller
Controller


