登録情報 データベース : EMDB / ID : EMD-16452タイトル Human spliceosomal PM5 C* complex map of the human spliceosomal PM5 C* complex 複合体 : human spliceosomal C* complex assembled on PM5 pre-mRNARNA : x 5種タンパク質・ペプチド : x 48種リガンド : x 6種 / / 機能・相同性 分子機能 ドメイン・相同性 構成要素
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / 生物種 Homo sapiens (ヒト) / human (ヒト)手法 / / 解像度 : 2.8 Å Dybkov O / Kastner B / Luehrmann R 資金援助 Organization Grant number 国 Max Planck Society
ジャーナル : Sci Adv / 年 : 2023タイトル : Regulation of 3' splice site selection after step 1 of splicing by spliceosomal C* proteins.著者: Olexandr Dybkov / Marco Preußner / Leyla El Ayoubi / Vivi-Yun Feng / Caroline Harnisch / Kilian Merz / Paula Leupold / Peter Yudichev / Dmitry E Agafonov / Cindy L Will / Cyrille Girard / ... 著者 : Olexandr Dybkov / Marco Preußner / Leyla El Ayoubi / Vivi-Yun Feng / Caroline Harnisch / Kilian Merz / Paula Leupold / Peter Yudichev / Dmitry E Agafonov / Cindy L Will / Cyrille Girard / Christian Dienemann / Henning Urlaub / Berthold Kastner / Florian Heyd / Reinhard Lührmann / 要旨 : Alternative precursor messenger RNA splicing is instrumental in expanding the proteome of higher eukaryotes, and changes in 3' splice site (3'ss) usage contribute to human disease. We demonstrate by ... Alternative precursor messenger RNA splicing is instrumental in expanding the proteome of higher eukaryotes, and changes in 3' splice site (3'ss) usage contribute to human disease. We demonstrate by small interfering RNA-mediated knockdowns, followed by RNA sequencing, that many proteins first recruited to human C* spliceosomes, which catalyze step 2 of splicing, regulate alternative splicing, including the selection of alternatively spliced NAGNAG 3'ss. Cryo-electron microscopy and protein cross-linking reveal the molecular architecture of these proteins in C* spliceosomes, providing mechanistic and structural insights into how they influence 3'ss usage. They further elucidate the path of the 3' region of the intron, allowing a structure-based model for how the C* spliceosome potentially scans for the proximal 3'ss. By combining biochemical and structural approaches with genome-wide functional analyses, our studies reveal widespread regulation of alternative 3'ss usage after step 1 of splicing and the likely mechanisms whereby C* proteins influence NAGNAG 3'ss choices. 履歴 登録 2023年1月12日 - ヘッダ(付随情報) 公開 2023年7月12日 - マップ公開 2023年7月12日 - 更新 2023年7月12日 - 現状 2023年7月12日 処理サイト : PDBe / 状態 : 公開
すべて表示 表示を減らす
 データを開く
データを開く 基本情報
基本情報
 マップデータ
マップデータ 試料
試料 キーワード
キーワード spliceosome (スプライセオソーム) /
spliceosome (スプライセオソーム) /  nucleus (細胞核) /
nucleus (細胞核) /  splicing
splicing 機能・相同性情報
機能・相同性情報 selenocysteine insertion sequence binding / exon-exon junction complex / NOSIP mediated eNOS trafficking / pre-mRNA 3'-splice site binding / granulocyte differentiation /
selenocysteine insertion sequence binding / exon-exon junction complex / NOSIP mediated eNOS trafficking / pre-mRNA 3'-splice site binding / granulocyte differentiation /  snRNP binding / post-mRNA release spliceosomal complex / U2 snRNP binding / regulation of retinoic acid receptor signaling pathway / negative regulation of toll-like receptor signaling pathway /
snRNP binding / post-mRNA release spliceosomal complex / U2 snRNP binding / regulation of retinoic acid receptor signaling pathway / negative regulation of toll-like receptor signaling pathway /  U7 snRNA binding / histone pre-mRNA DCP binding /
U7 snRNA binding / histone pre-mRNA DCP binding /  regulation of translation at postsynapse, modulating synaptic transmission / 3'-5' RNA helicase activity / U7 snRNP / generation of catalytic spliceosome for first transesterification step /
regulation of translation at postsynapse, modulating synaptic transmission / 3'-5' RNA helicase activity / U7 snRNP / generation of catalytic spliceosome for first transesterification step /  renal system process / negative regulation of excitatory postsynaptic potential / histone pre-mRNA 3'end processing complex / alternative mRNA splicing, via spliceosome / endonucleolytic cleavage of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / cis assembly of pre-catalytic spliceosome / regulation of vitamin D receptor signaling pathway / Z-decay: degradation of maternal mRNAs by zygotically expressed factors / negative regulation of catalytic activity / SLBP independent Processing of Histone Pre-mRNAs / SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs / negative regulation of interleukin-8 production /
renal system process / negative regulation of excitatory postsynaptic potential / histone pre-mRNA 3'end processing complex / alternative mRNA splicing, via spliceosome / endonucleolytic cleavage of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / cis assembly of pre-catalytic spliceosome / regulation of vitamin D receptor signaling pathway / Z-decay: degradation of maternal mRNAs by zygotically expressed factors / negative regulation of catalytic activity / SLBP independent Processing of Histone Pre-mRNAs / SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs / negative regulation of interleukin-8 production /  regulation of mRNA processing / negative regulation of lipopolysaccharide-mediated signaling pathway / Deadenylation of mRNA / spliceosome conformational change to release U4 (or U4atac) and U1 (or U11) / embryonic brain development / negative regulation of phosphorylation /
regulation of mRNA processing / negative regulation of lipopolysaccharide-mediated signaling pathway / Deadenylation of mRNA / spliceosome conformational change to release U4 (or U4atac) and U1 (or U11) / embryonic brain development / negative regulation of phosphorylation /  protein methylation / U12-type spliceosomal complex / methylosome / STING mediated induction of host immune responses / 7-methylguanosine cap hypermethylation / negative regulation of interferon-beta production / nuclear retinoic acid receptor binding / U2-type catalytic step 1 spliceosome / U1 snRNP binding / pICln-Sm protein complex / Prp19 complex /
protein methylation / U12-type spliceosomal complex / methylosome / STING mediated induction of host immune responses / 7-methylguanosine cap hypermethylation / negative regulation of interferon-beta production / nuclear retinoic acid receptor binding / U2-type catalytic step 1 spliceosome / U1 snRNP binding / pICln-Sm protein complex / Prp19 complex /  RNA splicing, via transesterification reactions /
RNA splicing, via transesterification reactions /  poly(A) binding / positive regulation of androgen receptor activity / M-decay: degradation of maternal mRNAs by maternally stored factors / RNA stem-loop binding / mRNA 3'-end processing / spliceosomal tri-snRNP complex /
poly(A) binding / positive regulation of androgen receptor activity / M-decay: degradation of maternal mRNAs by maternally stored factors / RNA stem-loop binding / mRNA 3'-end processing / spliceosomal tri-snRNP complex /  small nuclear ribonucleoprotein complex / P granule /
small nuclear ribonucleoprotein complex / P granule /  pre-mRNA binding / sno(s)RNA-containing ribonucleoprotein complex / ATP-dependent activity, acting on RNA / SMN-Sm protein complex / embryonic cranial skeleton morphogenesis / mRNA cis splicing, via spliceosome / U2-type spliceosomal complex /
pre-mRNA binding / sno(s)RNA-containing ribonucleoprotein complex / ATP-dependent activity, acting on RNA / SMN-Sm protein complex / embryonic cranial skeleton morphogenesis / mRNA cis splicing, via spliceosome / U2-type spliceosomal complex /  telomerase RNA binding / U2-type precatalytic spliceosome / IRF3-mediated induction of type I IFN / positive regulation of mRNA splicing, via spliceosome /
telomerase RNA binding / U2-type precatalytic spliceosome / IRF3-mediated induction of type I IFN / positive regulation of mRNA splicing, via spliceosome /  telomerase holoenzyme complex / C2H2 zinc finger domain binding /
telomerase holoenzyme complex / C2H2 zinc finger domain binding /  regulation of mRNA splicing, via spliceosome / U2-type prespliceosome assembly / U2-type catalytic step 2 spliceosome / commitment complex / positive regulation by host of viral transcription / U4 snRNP / positive regulation of vitamin D receptor signaling pathway / Transport of Mature mRNA derived from an Intron-Containing Transcript / Notch binding / U2 snRNP / nuclear vitamin D receptor binding /
regulation of mRNA splicing, via spliceosome / U2-type prespliceosome assembly / U2-type catalytic step 2 spliceosome / commitment complex / positive regulation by host of viral transcription / U4 snRNP / positive regulation of vitamin D receptor signaling pathway / Transport of Mature mRNA derived from an Intron-Containing Transcript / Notch binding / U2 snRNP / nuclear vitamin D receptor binding /  Regulation of gene expression in late stage (branching morphogenesis) pancreatic bud precursor cells / RNA Polymerase II Transcription Termination / RUNX3 regulates NOTCH signaling / U1 snRNP / positive regulation of alpha-beta T cell differentiation / NOTCH4 Intracellular Domain Regulates Transcription / U2-type prespliceosome / ubiquitin-ubiquitin ligase activity / nuclear-transcribed mRNA catabolic process, nonsense-mediated decay / K63-linked polyubiquitin modification-dependent protein binding / WD40-repeat domain binding / NOTCH3 Intracellular Domain Regulates Transcription
Regulation of gene expression in late stage (branching morphogenesis) pancreatic bud precursor cells / RNA Polymerase II Transcription Termination / RUNX3 regulates NOTCH signaling / U1 snRNP / positive regulation of alpha-beta T cell differentiation / NOTCH4 Intracellular Domain Regulates Transcription / U2-type prespliceosome / ubiquitin-ubiquitin ligase activity / nuclear-transcribed mRNA catabolic process, nonsense-mediated decay / K63-linked polyubiquitin modification-dependent protein binding / WD40-repeat domain binding / NOTCH3 Intracellular Domain Regulates Transcription
 Homo sapiens (ヒト) /
Homo sapiens (ヒト) / 
 human (ヒト)
human (ヒト) 単粒子再構成法 /
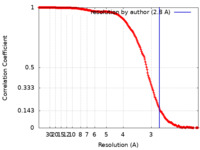
単粒子再構成法 /  クライオ電子顕微鏡法 / 解像度: 2.8 Å
クライオ電子顕微鏡法 / 解像度: 2.8 Å  データ登録者
データ登録者 ドイツ, 1件
ドイツ, 1件  引用
引用 ジャーナル: Sci Adv / 年: 2023
ジャーナル: Sci Adv / 年: 2023
 構造の表示
構造の表示 ダウンロードとリンク
ダウンロードとリンク emd_16452.map.gz
emd_16452.map.gz EMDBマップデータ形式
EMDBマップデータ形式 emd-16452-v30.xml
emd-16452-v30.xml emd-16452.xml
emd-16452.xml EMDBヘッダ
EMDBヘッダ emd_16452_fsc.xml
emd_16452_fsc.xml FSCデータファイル
FSCデータファイル emd_16452.png
emd_16452.png emd_16452_additional_1.map.gz
emd_16452_additional_1.map.gz emd_16452_half_map_1.map.gz
emd_16452_half_map_1.map.gz emd_16452_half_map_2.map.gz
emd_16452_half_map_2.map.gz http://ftp.pdbj.org/pub/emdb/structures/EMD-16452
http://ftp.pdbj.org/pub/emdb/structures/EMD-16452 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16452
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16452
 F&H 検索
F&H 検索 リンク
リンク EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource マップ
マップ ダウンロード / ファイル: emd_16452.map.gz / 形式: CCP4 / 大きさ: 744.3 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
ダウンロード / ファイル: emd_16452.map.gz / 形式: CCP4 / 大きさ: 744.3 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) 試料の構成要素
試料の構成要素 クライオ電子顕微鏡法
クライオ電子顕微鏡法 解析
解析 単粒子再構成法
単粒子再構成法 試料調製
試料調製 電子顕微鏡法
電子顕微鏡法 FIELD EMISSION GUN
FIELD EMISSION GUN Bright-field microscopy / Cs: 2.7 mm / 最大 デフォーカス(公称値): 3.0 µm / 最小 デフォーカス(公称値): 0.5 µm / 倍率(公称値): 81000
Bright-field microscopy / Cs: 2.7 mm / 最大 デフォーカス(公称値): 3.0 µm / 最小 デフォーカス(公称値): 0.5 µm / 倍率(公称値): 81000 
 画像解析
画像解析
 ムービー
ムービー コントローラー
コントローラー




































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)