[English] 日本語
 Yorodumi
Yorodumi- SASDBL2: MBP-PICK1 fusion (Maltose Binding Protein fused to Protein Intera... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: SASBDB / ID: SASDBL2 |
|---|---|
 Sample Sample | MBP-PICK1 fusion
|
| Function / homology |  Function and homology information Function and homology informationmembrane curvature sensor activity / glial cell development / neuronal ion channel clustering / cellular response to decreased oxygen levels / Trafficking of GluR2-containing AMPA receptors / Arp2/3 complex binding / regulation of Arp2/3 complex-mediated actin nucleation / negative regulation of Arp2/3 complex-mediated actin nucleation / : / dendritic spine organization ...membrane curvature sensor activity / glial cell development / neuronal ion channel clustering / cellular response to decreased oxygen levels / Trafficking of GluR2-containing AMPA receptors / Arp2/3 complex binding / regulation of Arp2/3 complex-mediated actin nucleation / negative regulation of Arp2/3 complex-mediated actin nucleation / : / dendritic spine organization / dendritic spine maintenance / long-term synaptic depression / receptor clustering / regulation of insulin secretion / positive regulation of receptor internalization / cellular response to glucose starvation / protein kinase C binding / Cell surface interactions at the vascular wall / trans-Golgi network membrane / intracellular protein transport / phospholipid binding / G protein-coupled receptor binding / actin filament binding / endocytic vesicle membrane / synaptic vesicle / presynaptic membrane / cytoskeleton / protein phosphorylation / neuron projection / postsynaptic density / signaling receptor binding / protein domain specific binding / synapse / perinuclear region of cytoplasm / Golgi apparatus / metal ion binding / identical protein binding / plasma membrane / cytoplasm / cytosol Similarity search - Function |
| Biological species |  Homo sapiens (human) Homo sapiens (human) |
 Citation Citation |  Journal: Mol Biol Cell / Year: 2015 Journal: Mol Biol Cell / Year: 2015Title: PICK1 is implicated in organelle motility in an Arp2/3 complex-independent manner. Authors: Yadaiah Madasu / Changsong Yang / Malgorzata Boczkowska / Kelley A Bethoney / Adam Zwolak / Grzegorz Rebowski / Tatyana Svitkina / Roberto Dominguez /  Abstract: PICK1 is a modular scaffold implicated in synaptic receptor trafficking. It features a PDZ domain, a BAR domain, and an acidic C-terminal tail (ACT). Analysis by small- angle x-ray scattering ...PICK1 is a modular scaffold implicated in synaptic receptor trafficking. It features a PDZ domain, a BAR domain, and an acidic C-terminal tail (ACT). Analysis by small- angle x-ray scattering suggests a structural model that places the receptor-binding site of the PDZ domain and membrane-binding surfaces of the BAR and PDZ domains adjacent to each other on the concave side of the banana-shaped PICK1 dimer. In the model, the ACT of one subunit of the dimer interacts with the PDZ and BAR domains of the other subunit, possibly accounting for autoinhibition. Consistently, full-length PICK1 shows diffuse cytoplasmic localization, but it clusters on vesicle-like structures that colocalize with the trans-Golgi network marker TGN38 upon deletion of either the ACT or PDZ domain. This localization is driven by the BAR domain. Live-cell imaging further reveals that PICK1-associated vesicles undergo fast, nondirectional motility in an F-actin-dependent manner, but deleting the ACT dramatically reduces vesicle speed. Thus the ACT links PICK1-associated vesicles to a motility factor, likely myosin, but, contrary to previous reports, PICK1 neither binds nor inhibits Arp2/3 complex. |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
-Data source
| SASBDB page |  SASDBL2 SASDBL2 |
|---|
-Related structure data
| Similar structure data |
|---|
- External links
External links
| Related items in Molecule of the Month |
|---|
-Models
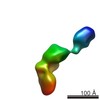
| Model #402 |  Type: dummy / Software: crysol / Radius of dummy atoms: 1.90 A / Chi-square value: 1.1881  Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
|---|
- Sample
Sample
 Sample Sample | Name: MBP-PICK1 fusion / Specimen concentration: 0.46-7.50 |
|---|---|
| Buffer | Name: 50 mM Tris / pH: 7.5 / Composition: 300 mM NaCl, 1 mM maltose, 1 mM EGTA, 2 mM DTT |
| Entity #266 | Name: MBP-PICK1 / Type: protein Description: Maltose Binding Protein fused to Protein Interacting with C kinase 1 Formula weight: 87.089 / Num. of mol.: 2 / Source: Homo sapiens / References: UniProt: Q9NRD5 Sequence: MKIEEGKLVI WINGDKGYNG LAEVGKKFEK DTGIKVTVEH PDKLEEKFPQ VAATGDGPDI IFWAHDRFGG YAQSGLLAEI TPDKAFQDKL YPFTWDAVRY NGKLIAYPIA VEALSLIYNK DLLPNPPKTW EEIPALDKEL KAKGKSALMF NLQEPYFTWP LIAADGGYAF ...Sequence: MKIEEGKLVI WINGDKGYNG LAEVGKKFEK DTGIKVTVEH PDKLEEKFPQ VAATGDGPDI IFWAHDRFGG YAQSGLLAEI TPDKAFQDKL YPFTWDAVRY NGKLIAYPIA VEALSLIYNK DLLPNPPKTW EEIPALDKEL KAKGKSALMF NLQEPYFTWP LIAADGGYAF KYENGKYDIK DVGVDNAGAK AGLTFLVDLI KNKHMNADTD YSIAEAAFNK GETAMTINGP WAWSNIDTSK VNYGVTVLPT FKGQPSKPFV GVLSAGINAA SPNKELAKEF LENYLLTDEG LEAVNKDKPL GAVALKSYEE ELAKDPRIAA TMENAQKGEI MPNIPQMSAF WYAVRTAVIN AASGRQTVDA ALAAAQTNAA AMFADLDYDI EEDKLGIPTV PGKVTLQKDA QNLIGISIGG GAQYCPCLYI VQVFDNTPAA LDGTVAAGDE ITGVNGRSIK GKTKVEVAKM IQEVKGEVTI HYNKLQADPK QGMSLDIVLK KVKHRLVENM SSGTADALGL SRAILCNDGL VKRLEELERT AELYKGMTEH TKNLLRAFYE LSQTHRAFGD VFSVIGVREP QPAASEAFVK FADAHRSIEK FGIRLLKTIK PMLTDLNTYL NKAIPDTRLT IKKYLDVKFE YLSYCLKVKE MDDEEYSCIA LGEPLYRVST GNYEYRLILR CRQEARARFS QMRKDVLEKM ELLDQKHVQD IVFQLQRLVS TMSKYYNDCY AVLRDADVFP IEVDLAHTTL AYGLNQEEFT DGEEEEEEED TAAGEPSRDT RGAAGPLDKG GSWCDS |
-Experimental information
| Beam | Instrument name: Cornell High Energy Synchrotron Source (CHESS) G1 City: Ithaca, NY / 国: USA  / Type of source: X-ray synchrotron / Type of source: X-ray synchrotron | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: Finger Lakes CCD / Type: CCD | ||||||||||||||||||
| Scan |
| ||||||||||||||||||
| Distance distribution function P(R) |
| ||||||||||||||||||
| Result | Comments: This dataset was collected at a concentration of 3.75 mg/ml.
|
 Movie
Movie Controller
Controller