[English] 日本語
 Yorodumi
Yorodumi- PDB-8rm6: Crystal Structure of Human Androgen Receptor DNA Binding Domain B... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8rm6 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal Structure of Human Androgen Receptor DNA Binding Domain Bound to its Response Element: C3(1)ARE | ||||||
 Components Components |
| ||||||
 Keywords Keywords | DNA BINDING PROTEIN / androgen receptor / androgen response elements / dbd binding domain-response element complex | ||||||
| Function / homology |  Function and homology information Function and homology informationmale somatic sex determination / prostate induction / lateral sprouting involved in mammary gland duct morphogenesis / male genitalia morphogenesis / regulation of developmental growth / POU domain binding / positive regulation of integrin biosynthetic process / tertiary branching involved in mammary gland duct morphogenesis / animal organ formation / androgen binding ...male somatic sex determination / prostate induction / lateral sprouting involved in mammary gland duct morphogenesis / male genitalia morphogenesis / regulation of developmental growth / POU domain binding / positive regulation of integrin biosynthetic process / tertiary branching involved in mammary gland duct morphogenesis / animal organ formation / androgen binding / cellular response to testosterone stimulus / regulation of systemic arterial blood pressure / Leydig cell differentiation / epithelial cell differentiation involved in prostate gland development / positive regulation of epithelial cell proliferation involved in prostate gland development / prostate gland epithelium morphogenesis / prostate gland growth / epithelial cell morphogenesis / membraneless organelle assembly / RNA polymerase II general transcription initiation factor binding / positive regulation of insulin-like growth factor receptor signaling pathway / positive regulation of transcription by RNA polymerase III / morphogenesis of an epithelial fold / cellular response to steroid hormone stimulus / positive regulation of intracellular estrogen receptor signaling pathway / androgen receptor signaling pathway / seminiferous tubule development / RUNX2 regulates osteoblast differentiation / nuclear steroid receptor activity / mammary gland alveolus development / cellular response to estrogen stimulus / estrogen response element binding / nuclear receptor-mediated steroid hormone signaling pathway / single fertilization / RNA polymerase II core promoter sequence-specific DNA binding / regulation of protein localization to plasma membrane / estrogen receptor signaling pathway / intracellular receptor signaling pathway / steroid binding / insulin-like growth factor receptor signaling pathway / HSP90 chaperone cycle for steroid hormone receptors (SHR) in the presence of ligand / epithelial cell proliferation / negative regulation of extrinsic apoptotic signaling pathway / RNA polymerase II transcription regulatory region sequence-specific DNA binding / SUMOylation of intracellular receptors / positive regulation of cell differentiation / molecular condensate scaffold activity / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / beta-catenin binding / Nuclear Receptor transcription pathway / positive regulation of miRNA transcription / male gonad development / transcription coactivator binding / multicellular organism growth / nuclear receptor activity / negative regulation of epithelial cell proliferation / cell-cell signaling / MAPK cascade / ATPase binding / DNA-binding transcription activator activity, RNA polymerase II-specific / spermatogenesis / in utero embryonic development / molecular adaptor activity / transcription by RNA polymerase II / RNA polymerase II-specific DNA-binding transcription factor binding / DNA-binding transcription factor activity, RNA polymerase II-specific / positive regulation of MAPK cascade / transcription cis-regulatory region binding / Ub-specific processing proteases / nuclear speck / RNA polymerase II cis-regulatory region sequence-specific DNA binding / DNA-binding transcription factor activity / signaling receptor binding / negative regulation of cell population proliferation / positive regulation of cell population proliferation / chromatin binding / positive regulation of gene expression / positive regulation of DNA-templated transcription / chromatin / enzyme binding / negative regulation of transcription by RNA polymerase II / signal transduction / positive regulation of transcription by RNA polymerase II / protein-containing complex / zinc ion binding / nucleoplasm / nucleus / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.05 Å MOLECULAR REPLACEMENT / Resolution: 2.05 Å | ||||||
 Authors Authors | Lee, X.Y. / Helsen, C. / Van Eynde, W. / Voet, A. / Claessens, F. | ||||||
| Funding support |  Belgium, 1items Belgium, 1items
| ||||||
 Citation Citation |  Journal: J.Steroid Biochem.Mol.Biol. / Year: 2024 Journal: J.Steroid Biochem.Mol.Biol. / Year: 2024Title: Structural mechanism underlying variations in DNA binding by the androgen receptor. Authors: Lee, X.Y. / Van Eynde, W. / Helsen, C. / Willems, H. / Peperstraete, K. / De Block, S. / Voet, A. / Claessens, F. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8rm6.cif.gz 8rm6.cif.gz | 65 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8rm6.ent.gz pdb8rm6.ent.gz | 43.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8rm6.json.gz 8rm6.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/rm/8rm6 https://data.pdbj.org/pub/pdb/validation_reports/rm/8rm6 ftp://data.pdbj.org/pub/pdb/validation_reports/rm/8rm6 ftp://data.pdbj.org/pub/pdb/validation_reports/rm/8rm6 | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 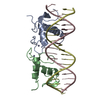 8rm7C C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||||
| Unit cell |
| ||||||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 8049.417 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: AR, DHTR, NR3C4 / Production host: Homo sapiens (human) / Gene: AR, DHTR, NR3C4 / Production host:  #2: DNA chain | | Mass: 5536.599 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:  #3: DNA chain | | Mass: 5483.593 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:  #4: Chemical | ChemComp-ZN / #5: Water | ChemComp-HOH / | Has ligand of interest | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.5 Å3/Da / Density % sol: 64.82 % |
|---|---|
| Crystal grow | Temperature: 293.15 K / Method: vapor diffusion, sitting drop / Details: 0.1M sodium hepes pH7.5,15% (w/v) peg 20000 |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID23-2 / Wavelength: 0.8731 Å / Beamline: ID23-2 / Wavelength: 0.8731 Å |
| Detector | Type: DECTRIS PILATUS3 X 2M / Detector: PIXEL / Date: Feb 14, 2021 |
| Radiation | Monochromator: M / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.8731 Å / Relative weight: 1 |
| Reflection | Resolution: 2.05→49.35 Å / Num. obs: 26537 / % possible obs: 99.9 % / Redundancy: 51.3 % / Biso Wilson estimate: 41.5 Å2 / CC1/2: 0.999 / Rmerge(I) obs: 0.239 / Rpim(I) all: 0.033 / Rrim(I) all: 0.242 / Net I/σ(I): 19.9 |
| Reflection shell | Resolution: 2.05→2.11 Å / Redundancy: 42.8 % / Rmerge(I) obs: 2.509 / Mean I/σ(I) obs: 2.5 / Num. unique obs: 1972 / CC1/2: 0.821 / Rpim(I) all: 0.386 / Rrim(I) all: 2.539 / % possible all: 99.4 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 2.05→49.35 Å / SU ML: 0.28 / Cross valid method: FREE R-VALUE / σ(F): 1.36 / Phase error: 25.45 / Stereochemistry target values: ML MOLECULAR REPLACEMENT / Resolution: 2.05→49.35 Å / SU ML: 0.28 / Cross valid method: FREE R-VALUE / σ(F): 1.36 / Phase error: 25.45 / Stereochemistry target values: ML
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.05→49.35 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller


 PDBj
PDBj












































