+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: PDB / ID: 7oqt | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | G-quadruplex structure of the C. elegans telomeric repeat: A two tetrads basket type conformation stabilised by a Hoogsteen C-T base-pair | ||||||||||||||||||||||||||||
 要素 要素 | DNA (5'-D(* キーワード キーワードDNA / G-quadruplex / C. elegans telomere | 機能・相同性 | : / DNA / DNA (> 10) |  機能・相同性情報 機能・相同性情報生物種 |  手法 | 溶液NMR / molecular dynamics |  データ登録者 データ登録者Marquevielle, J. / De Rache, A. / Amrane, S. | 資金援助 | 1件 |
 引用 引用 ジャーナル: Nucleic Acids Res. / 年: 2022 ジャーナル: Nucleic Acids Res. / 年: 2022タイトル: G-quadruplex structure of the C. elegans telomeric repeat: a two tetrads basket type conformation stabilized by a non-canonical C-T base-pair. 著者: Marquevielle, J. / De Rache, A. / Vialet, B. / Morvan, E. / Mergny, J.L. / Amrane, S. 履歴 |
|
- 構造の表示
構造の表示
| 構造ビューア | 分子:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- ダウンロードとリンク
ダウンロードとリンク
- ダウンロード
ダウンロード
| PDBx/mmCIF形式 |  7oqt.cif.gz 7oqt.cif.gz | 132.9 KB | 表示 |  PDBx/mmCIF形式 PDBx/mmCIF形式 |
|---|---|---|---|---|
| PDB形式 |  pdb7oqt.ent.gz pdb7oqt.ent.gz | 105.9 KB | 表示 |  PDB形式 PDB形式 |
| PDBx/mmJSON形式 |  7oqt.json.gz 7oqt.json.gz | ツリー表示 |  PDBx/mmJSON形式 PDBx/mmJSON形式 | |
| その他 |  その他のダウンロード その他のダウンロード |
-検証レポート
| 文書・要旨 |  7oqt_validation.pdf.gz 7oqt_validation.pdf.gz | 409.9 KB | 表示 |  wwPDB検証レポート wwPDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  7oqt_full_validation.pdf.gz 7oqt_full_validation.pdf.gz | 588.9 KB | 表示 | |
| XML形式データ |  7oqt_validation.xml.gz 7oqt_validation.xml.gz | 38.8 KB | 表示 | |
| CIF形式データ |  7oqt_validation.cif.gz 7oqt_validation.cif.gz | 31.2 KB | 表示 | |
| アーカイブディレクトリ |  https://data.pdbj.org/pub/pdb/validation_reports/oq/7oqt https://data.pdbj.org/pub/pdb/validation_reports/oq/7oqt ftp://data.pdbj.org/pub/pdb/validation_reports/oq/7oqt ftp://data.pdbj.org/pub/pdb/validation_reports/oq/7oqt | HTTPS FTP |
-関連構造データ
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
|---|---|
| その他のデータベース |
|
- リンク
リンク
- 集合体
集合体
| 登録構造単位 | 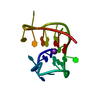
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR アンサンブル |
|
- 要素
要素
| #1: DNA鎖 | 分子量: 6221.012 Da / 分子数: 1 / 由来タイプ: 合成 / 由来: (合成)  |
|---|---|
| #2: 化合物 | ChemComp-K / |
| 研究の焦点であるリガンドがあるか | Y |
-実験情報
-実験
| 実験 | 手法: 溶液NMR | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR実験 |
|
- 試料調製
試料調製
| 詳細 | タイプ: solution / 内容: 2.4 mM 1H Asc20, 90% H2O/10% D2O / Label: Asc20 / 溶媒系: 90% H2O/10% D2O |
|---|---|
| 試料 | 濃度: 2.4 mM / 構成要素: Asc20 / Isotopic labeling: 1H |
| 試料状態 | イオン強度: 90 mM / Label: NMR buffer / pH: 6.9 / 圧: 1 atm / 温度: 288 K |
-NMR測定
| NMRスペクトロメーター | タイプ: Bruker AVANCE III / 製造業者: Bruker / モデル: AVANCE III / 磁場強度: 700 MHz |
|---|
- 解析
解析
| NMR software |
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 精密化 | 手法: molecular dynamics / ソフトェア番号: 4 | |||||||||||||||
| 代表構造 | 選択基準: lowest energy | |||||||||||||||
| NMRアンサンブル | コンフォーマー選択の基準: structures with the lowest energy 計算したコンフォーマーの数: 750 / 登録したコンフォーマーの数: 10 |
 ムービー
ムービー コントローラー
コントローラー



 PDBj
PDBj








































 HSQC
HSQC