+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7ea1 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal Structure of Spindlin1 bound to SPINDOC Docpep2 | ||||||
 Components Components |
| ||||||
 Keywords Keywords | TRANSCRIPTION / Spin/Ssty repeat | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of protein ADP-ribosylation / transposable element silencing by piRNA-mediated DNA methylation / gamete generation / histone H3K4me3 reader activity / : / histone reader activity / rRNA transcription / site of DNA damage / positive regulation of Wnt signaling pathway / protein localization to chromatin ...regulation of protein ADP-ribosylation / transposable element silencing by piRNA-mediated DNA methylation / gamete generation / histone H3K4me3 reader activity / : / histone reader activity / rRNA transcription / site of DNA damage / positive regulation of Wnt signaling pathway / protein localization to chromatin / meiotic cell cycle / spindle / Wnt signaling pathway / nuclear membrane / negative regulation of DNA-templated transcription / DNA damage response / regulation of DNA-templated transcription / chromatin / positive regulation of DNA-templated transcription / nucleolus / nucleoplasm / nucleus / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.7 Å MOLECULAR REPLACEMENT / Resolution: 2.7 Å | ||||||
 Authors Authors | Zhao, F. / Li, H. | ||||||
 Citation Citation |  Journal: to be published Journal: to be publishedTitle: Molecular basis for SPINDOC-Spindlin1 engagement and its role in transcriptional inhibition Authors: Zhao, F. / Li, H. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7ea1.cif.gz 7ea1.cif.gz | 176.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7ea1.ent.gz pdb7ea1.ent.gz | 139.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7ea1.json.gz 7ea1.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  7ea1_validation.pdf.gz 7ea1_validation.pdf.gz | 452.6 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  7ea1_full_validation.pdf.gz 7ea1_full_validation.pdf.gz | 463.8 KB | Display | |
| Data in XML |  7ea1_validation.xml.gz 7ea1_validation.xml.gz | 16.8 KB | Display | |
| Data in CIF |  7ea1_validation.cif.gz 7ea1_validation.cif.gz | 21.7 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ea/7ea1 https://data.pdbj.org/pub/pdb/validation_reports/ea/7ea1 ftp://data.pdbj.org/pub/pdb/validation_reports/ea/7ea1 ftp://data.pdbj.org/pub/pdb/validation_reports/ea/7ea1 | HTTPS FTP |
-Related structure data
| Related structure data | 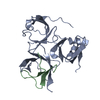 7e9mC  2ns2S S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 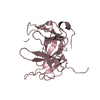
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 24549.670 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: SPIN1, OCR, SPIN / Plasmid: pGEX-6P-1 / Production host: Homo sapiens (human) / Gene: SPIN1, OCR, SPIN / Plasmid: pGEX-6P-1 / Production host:  #2: Protein/peptide | Mass: 1476.836 Da / Num. of mol.: 2 / Source method: obtained synthetically / Source: (synth.)  Homo sapiens (human) / References: UniProt: Q9BUA3 Homo sapiens (human) / References: UniProt: Q9BUA3#3: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.33 Å3/Da / Density % sol: 47.15 % / Mosaicity: 1.069 ° / Mosaicity esd: 0.011 ° |
|---|---|
| Crystal grow | Temperature: 291 K / Method: vapor diffusion / pH: 6.5 Details: 0.1M Sodium chloride, 0.1M BIS-TRIS pH 6.5, 1.5M Ammonium sulfate |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRF SSRF  / Beamline: BL17U / Wavelength: 0.97853 Å / Beamline: BL17U / Wavelength: 0.97853 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Type: ADSC QUANTUM 315r / Detector: CCD / Date: Feb 3, 2018 / Details: mirrors | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Monochromator: double crystal, Si(111) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 0.97853 Å / Relative weight: 1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 2.689→50 Å / Num. obs: 14134 / % possible obs: 99.8 % / Redundancy: 6.4 % / Rmerge(I) obs: 0.103 / Rpim(I) all: 0.045 / Rrim(I) all: 0.113 / Χ2: 1.085 / Net I/σ(I): 5.1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1
|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 2NS2 Resolution: 2.7→40.363 Å / SU ML: 0.43 / Cross valid method: THROUGHOUT / σ(F): 1.37 / Phase error: 34.46 / Stereochemistry target values: ML
| ||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 234.11 Å2 / Biso mean: 100.4159 Å2 / Biso min: 35.4 Å2 | ||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 2.7→40.363 Å
| ||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Refine-ID: X-RAY DIFFRACTION / Rfactor Rfree error: 0
| ||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: -0.525 Å / Origin y: 9.626 Å / Origin z: -6.152 Å
| ||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller



 PDBj
PDBj