[English] 日本語
 Yorodumi
Yorodumi- PDB-6q6m: RORCVAR2 (RORGT, 264-499) IN COMPLEX WITH COMPOUND 1: Identificat... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6q6m | ||||||
|---|---|---|---|---|---|---|---|
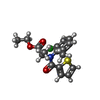
| Title | RORCVAR2 (RORGT, 264-499) IN COMPLEX WITH COMPOUND 1: Identification of N-aryl imidazoles as potent and selective RORgt inhibitors | ||||||
 Components Components | Nuclear receptor ROR-gamma | ||||||
 Keywords Keywords | TRANSCRIPTION / nuclear hormone receptor / ligand-binding domain / inverse agonist | ||||||
| Function / homology |  Function and homology information Function and homology informationtolerance induction in gut-associated lymphoid tissue / T-helper 17 cell differentiation / cellular response to sterol / regulation of steroid metabolic process / ligand-modulated transcription factor activity / Peyer's patch development / regulatory T cell differentiation / T-helper cell differentiation / positive regulation of circadian rhythm / RUNX3 Regulates Immune Response and Cell Migration ...tolerance induction in gut-associated lymphoid tissue / T-helper 17 cell differentiation / cellular response to sterol / regulation of steroid metabolic process / ligand-modulated transcription factor activity / Peyer's patch development / regulatory T cell differentiation / T-helper cell differentiation / positive regulation of circadian rhythm / RUNX3 Regulates Immune Response and Cell Migration / oxysterol binding / negative regulation of thymocyte apoptotic process / regulation of fat cell differentiation / Phosphorylated BMAL1:CLOCK (ARNTL:CLOCK) activates expression of core clock genes / regulation of glucose metabolic process / lymph node development / adipose tissue development / xenobiotic metabolic process / RORA,B,C and NR1D1 (REV-ERBA) regulate gene expression / Expression of BMAL (ARNTL), CLOCK, and NPAS2 / circadian regulation of gene expression / Nuclear Receptor transcription pathway / DNA-binding transcription repressor activity, RNA polymerase II-specific / nuclear receptor activity / sequence-specific double-stranded DNA binding / Interleukin-4 and Interleukin-13 signaling / DNA-binding transcription factor activity, RNA polymerase II-specific / nuclear body / RNA polymerase II cis-regulatory region sequence-specific DNA binding / DNA-binding transcription factor activity / regulation of transcription by RNA polymerase II / positive regulation of DNA-templated transcription / chromatin / negative regulation of transcription by RNA polymerase II / zinc ion binding / nucleoplasm / nucleus Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 2.35 Å molecular replacement / Resolution: 2.35 Å | ||||||
 Authors Authors | Kallen, J. | ||||||
 Citation Citation |  Journal: J.Med.Chem. / Year: 2019 Journal: J.Med.Chem. / Year: 2019Title: Structure-Based and Property-Driven Optimization ofN-Aryl Imidazoles toward Potent and Selective Oral ROR gamma t Inhibitors. Authors: Hoegenauer, K. / Kallen, J. / Jimenez-Nunez, E. / Strang, R. / Ertl, P. / Cooke, N.G. / Hintermann, S. / Voegtle, M. / Betschart, C. / McKay, D.J.J. / Wagner, J. / Ottl, J. / Beerli, C. / ...Authors: Hoegenauer, K. / Kallen, J. / Jimenez-Nunez, E. / Strang, R. / Ertl, P. / Cooke, N.G. / Hintermann, S. / Voegtle, M. / Betschart, C. / McKay, D.J.J. / Wagner, J. / Ottl, J. / Beerli, C. / Billich, A. / Dawson, J. / Kaupmann, K. / Streiff, M. / Gobeau, N. / Harlfinger, S. / Stringer, R. / Guntermann, C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6q6m.cif.gz 6q6m.cif.gz | 62.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6q6m.ent.gz pdb6q6m.ent.gz | 43.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6q6m.json.gz 6q6m.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/q6/6q6m https://data.pdbj.org/pub/pdb/validation_reports/q6/6q6m ftp://data.pdbj.org/pub/pdb/validation_reports/q6/6q6m ftp://data.pdbj.org/pub/pdb/validation_reports/q6/6q6m | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  6q6oC  6q7aC  6q7hC  5ntiS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 27629.943 Da / Num. of mol.: 1 / Fragment: C-terminal domain, ligand binding domain / Mutation: C455S Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: RORC, NR1F3, RORG, RZRG / Production host: Homo sapiens (human) / Gene: RORC, NR1F3, RORG, RZRG / Production host:  |
|---|---|
| #2: Chemical | ChemComp-HJW / |
| #3: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.5 Å3/Da / Density % sol: 64.89 % / Mosaicity: 0.484 ° |
|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 8.2 Details: 0.5% PEG MME 5k (w/v), 0.6 M KNa Tartrate 0.1M TRIS |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SLS SLS  / Beamline: X10SA / Wavelength: 0.9794 Å / Beamline: X10SA / Wavelength: 0.9794 Å |
| Detector | Type: MARMOSAIC 225 mm CCD / Detector: CCD / Date: Nov 7, 2008 |
| Radiation | Monochromator: SI 111 channel / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9794 Å / Relative weight: 1 |
| Reflection | Resolution: 2.35→20 Å / Num. obs: 17081 / % possible obs: 99.9 % / Redundancy: 13.7 % / Rmerge(I) obs: 0.068 / Χ2: 1.065 / Net I/σ(I): 45.26 |
| Reflection shell | Resolution: 2.35→2.43 Å / Redundancy: 14.2 % / Rmerge(I) obs: 0.327 / Mean I/σ(I) obs: 5.43 / Num. unique obs: 1859 / Χ2: 2.091 / % possible all: 100 |
-Phasing
| Phasing | Method:  molecular replacement molecular replacement |
|---|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 5NTI Resolution: 2.35→20 Å / Cor.coef. Fo:Fc: 0.914 / Cor.coef. Fo:Fc free: 0.91 / SU B: 6.725 / SU ML: 0.163 / SU R Cruickshank DPI: 0.2828 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.283 / ESU R Free: 0.224 Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS U VALUES : REFINED INDIVIDUALLY
| |||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å | |||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 79.74 Å2 / Biso mean: 48.133 Å2 / Biso min: 22.99 Å2
| |||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 2.35→20 Å
| |||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.35→2.41 Å / Rfactor Rfree error: 0 / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller











 PDBj
PDBj