+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5gwq | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of two TTTA repeats | ||||||||||||||||||||||||||||
 Components Components | DNA (5'-D(* Keywords KeywordsDNA / Minidumbbell / type II loop | Function / homology | DNA |  Function and homology information Function and homology informationBiological species |  Method | SOLUTION NMR / restrained molecular dynamics |  Authors AuthorsGuo, P. / Lam, S.L. | Funding support | |  Hong Kong, 1items Hong Kong, 1items
 Citation Citation Journal: J.Am.Chem.Soc. / Year: 2016 Journal: J.Am.Chem.Soc. / Year: 2016Title: Minidumbbell: A New Form of Native DNA Structure Authors: Guo, P. / Lam, S.L. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5gwq.cif.gz 5gwq.cif.gz | 97.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5gwq.ent.gz pdb5gwq.ent.gz | 66 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5gwq.json.gz 5gwq.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/gw/5gwq https://data.pdbj.org/pub/pdb/validation_reports/gw/5gwq ftp://data.pdbj.org/pub/pdb/validation_reports/gw/5gwq ftp://data.pdbj.org/pub/pdb/validation_reports/gw/5gwq | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 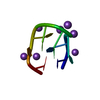
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: DNA chain | Mass: 2406.613 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.)  |
|---|---|
| #2: Chemical | ChemComp-NA / |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Type: solution Contents: 1.0 mM DNA (5'-D(*TP*TP*TP*AP*TP*TP*TP*A)-3'), 10 mM sodium phosphate, 0.1 mM DSS, 100% D2O Label: 1H / Solvent system: 100% D2O | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||
| Sample conditions | Ionic strength: 0.02 M / Label: conditions_1 / pH: 7 / Pressure: 1 atm / Temperature: 278 K |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: restrained molecular dynamics / Software ordinal: 2 | |||||||||
| NMR representative | Selection criteria: lowest energy | |||||||||
| NMR ensemble | Conformer selection criteria: structures with the least restraint violations Conformers calculated total number: 1000 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller






 PDBj
PDBj



 Amber
Amber