+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5a9p | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
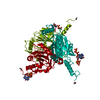
| Title | Crystal structure of Operophtera brumata CPV18 polyhedra | |||||||||
 Components Components | POLYHEDRIN | |||||||||
 Keywords Keywords | VIRAL PROTEIN / INSECT VIRUS OCCLUSION BODY / MICROCRYSTAL | |||||||||
| Function / homology | Cypovirus polyhedrin, Cypovirus 1 type / Cypovirus polyhedrin protein / ATP binding / metal ion binding / ADENOSINE-5'-TRIPHOSPHATE / GUANOSINE-5'-TRIPHOSPHATE / Polyhedrin Function and homology information Function and homology information | |||||||||
| Biological species |  OPEROPHTERA BRUMATA CYPOVIRUS 18 OPEROPHTERA BRUMATA CYPOVIRUS 18 | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.476 Å MOLECULAR REPLACEMENT / Resolution: 1.476 Å | |||||||||
 Authors Authors | Ji, X. / Axford, D. / Owen, R. / Evans, G. / Ginn, H.M. / Sutton, G. / Stuart, D.I. | |||||||||
 Citation Citation |  Journal: J.Struct.Biol. / Year: 2015 Journal: J.Struct.Biol. / Year: 2015Title: Polyhedra Structures and the Evolution of the Insect Viruses. Authors: Ji, X. / Axford, D. / Owen, R. / Evans, G. / Ginn, H.M. / Sutton, G. / Stuart, D.I. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5a9p.cif.gz 5a9p.cif.gz | 150.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5a9p.ent.gz pdb5a9p.ent.gz | 119.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5a9p.json.gz 5a9p.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/a9/5a9p https://data.pdbj.org/pub/pdb/validation_reports/a9/5a9p ftp://data.pdbj.org/pub/pdb/validation_reports/a9/5a9p ftp://data.pdbj.org/pub/pdb/validation_reports/a9/5a9p | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  5a8sC  5a8tC  5a8uC  5a8vC  5a96C  5a98C  5a99C  5a9aC  5a9bC  5a9cC  2oh6S C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| |||||||||||||||
| Unit cell |
| |||||||||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 28406.531 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  OPEROPHTERA BRUMATA CYPOVIRUS 18 / Cell line (production host): SF9 / Production host: OPEROPHTERA BRUMATA CYPOVIRUS 18 / Cell line (production host): SF9 / Production host:  |
|---|---|
| #2: Chemical | ChemComp-ATP / |
| #3: Chemical | ChemComp-GTP / |
| #4: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 21 X-RAY DIFFRACTION / Number of used crystals: 21 |
|---|
- Sample preparation
Sample preparation
| Crystal | Description: NONE |
|---|---|
| Crystal grow | pH: 7.5 / Details: PH 7.5 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Diamond Diamond  / Beamline: I24 / Wavelength: 0.979 / Beamline: I24 / Wavelength: 0.979 |
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.979 Å / Relative weight: 1 |
| Reflection | Resolution: 1.48→72.7 Å / Num. obs: 30415 / % possible obs: 100 % / Observed criterion σ(I): 1 / Redundancy: 12.9 % / Rmerge(I) obs: 0.32 / Net I/σ(I): 7.1 |
| Reflection shell | Resolution: 1.48→1.51 Å / Redundancy: 8.6 % / Mean I/σ(I) obs: 1.5 / % possible all: 99.8 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 2OH6 Resolution: 1.476→72.669 Å / SU ML: 0.13 / σ(F): 1.34 / Phase error: 13.71 / Stereochemistry target values: ML
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.476→72.669 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: 34.1141 Å / Origin y: 21.1181 Å / Origin z: 8.7019 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Selection details: ALL |
 Movie
Movie Controller
Controller








 PDBj
PDBj




