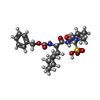
Entry Database : PDB / ID : 4xbdTitle 1.45A resolution structure of Norovirus 3CL protease complex with a covalently bound dipeptidyl inhibitor (1R,2S)-2-({N-[(benzyloxy)carbonyl]-3-cyclohexyl-L-alanyl}amino)-1-hydroxy-3-[(3S)-2-oxopyrrolidin-3-yl]propane-1-sulfonic acid (Orthorhombic P Form) 3C-LIKE PROTEASE Keywords / / / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Method / / / / Resolution : 1.45 Å Authors Lovell, S. / Battaile, K.P. / Mehzabeen, N. / Kankanamalage, A.C.G. / Kim, Y. / Weerawarna, P.M. / Uy, R.A.Z. / Damalanka, V.C. / Mandadapu, S.R. / Alliston, K.R. ...Lovell, S. / Battaile, K.P. / Mehzabeen, N. / Kankanamalage, A.C.G. / Kim, Y. / Weerawarna, P.M. / Uy, R.A.Z. / Damalanka, V.C. / Mandadapu, S.R. / Alliston, K.R. / Groutas, W.C. / Chang, K.-O. Journal : J.Med.Chem. / Year : 2015Title : Structure-Guided Design and Optimization of Dipeptidyl Inhibitors of Norovirus 3CL Protease. Structure-Activity Relationships and Biochemical, X-ray Crystallographic, Cell-Based, and In Vivo Studies.Authors : Galasiti Kankanamalage, A.C. / Kim, Y. / Weerawarna, P.M. / Uy, R.A. / Damalanka, V.C. / Mandadapu, S.R. / Alliston, K.R. / Mehzabeen, N. / Battaile, K.P. / Lovell, S. / Chang, K.O. / Groutas, W.C. History Deposition Dec 16, 2014 Deposition site / Processing site Revision 1.0 Mar 25, 2015 Provider / Type Revision 1.1 Apr 22, 2015 Group Revision 1.2 Nov 22, 2017 Group Advisory / Database references ... Advisory / Database references / Derived calculations / Refinement description / Source and taxonomy Category citation / entity_src_gen ... citation / entity_src_gen / pdbx_struct_assembly / pdbx_struct_assembly_gen / pdbx_struct_assembly_prop / pdbx_struct_oper_list / pdbx_unobs_or_zero_occ_atoms / software Item _citation.journal_id_CSD / _entity_src_gen.pdbx_alt_source_flag ... _citation.journal_id_CSD / _entity_src_gen.pdbx_alt_source_flag / _pdbx_struct_assembly.oligomeric_details / _pdbx_struct_assembly_gen.asym_id_list / _pdbx_struct_assembly_prop.type / _pdbx_struct_assembly_prop.value / _pdbx_struct_oper_list.symmetry_operation / _software.classification Revision 1.3 Nov 13, 2024 Group Data collection / Database references ... Data collection / Database references / Derived calculations / Structure summary Category chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_entry_details / pdbx_modification_feature / struct_conn Item / _database_2.pdbx_database_accession / _struct_conn.pdbx_leaving_atom_flag
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT /
MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 1.45 Å
molecular replacement / Resolution: 1.45 Å  Authors
Authors Citation
Citation Journal: J.Med.Chem. / Year: 2015
Journal: J.Med.Chem. / Year: 2015 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 4xbd.cif.gz
4xbd.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb4xbd.ent.gz
pdb4xbd.ent.gz PDB format
PDB format 4xbd.json.gz
4xbd.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/xb/4xbd
https://data.pdbj.org/pub/pdb/validation_reports/xb/4xbd ftp://data.pdbj.org/pub/pdb/validation_reports/xb/4xbd
ftp://data.pdbj.org/pub/pdb/validation_reports/xb/4xbd Links
Links Assembly
Assembly


 Components
Components Norwalk virus (strain GI/Human/United States/Norwalk/1968)
Norwalk virus (strain GI/Human/United States/Norwalk/1968)
 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  APS
APS  / Beamline: 17-ID / Wavelength: 1 Å
/ Beamline: 17-ID / Wavelength: 1 Å molecular replacement
molecular replacement Processing
Processing MOLECULAR REPLACEMENT / Resolution: 1.45→35.654 Å / SU ML: 0.15 / Cross valid method: THROUGHOUT / σ(F): 1.84 / Phase error: 18.21 / Stereochemistry target values: ML
MOLECULAR REPLACEMENT / Resolution: 1.45→35.654 Å / SU ML: 0.15 / Cross valid method: THROUGHOUT / σ(F): 1.84 / Phase error: 18.21 / Stereochemistry target values: ML Movie
Movie Controller
Controller













 PDBj
PDBj