+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4v3g | ||||||
|---|---|---|---|---|---|---|---|
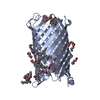
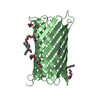
| Title | Crystal structure of CymA from Klebsiella oxytoca | ||||||
 Components Components | CYMA PROTEIN | ||||||
 Keywords Keywords | TRANSPORT PROTEIN / OUTER MEMBRANE CHANNEL CYCLODEXTRIN TRANSPORT BETA BARREL MONOMER | ||||||
| Function / homology | Cyclodextrin porin CymA / Putative cyclodextrin porin / import into cell / Outer membrane protein/outer membrane enzyme PagP, beta-barrel / CymA protein Function and homology information Function and homology information | ||||||
| Biological species |  KLEBSIELLA OXYTOCA (bacteria) KLEBSIELLA OXYTOCA (bacteria) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  SAD / Resolution: 2.513 Å SAD / Resolution: 2.513 Å | ||||||
 Authors Authors | van den Berg, B. / Bhamidimarri, S.P. / Kleinekathoefer, U. / Winterhalter, M. | ||||||
 Citation Citation |  Journal: Proc. Natl. Acad. Sci. U.S.A. / Year: 2015 Journal: Proc. Natl. Acad. Sci. U.S.A. / Year: 2015Title: Outer-membrane translocation of bulky small molecules by passive diffusion. Authors: van den Berg, B. / Prathyusha Bhamidimarri, S. / Dahyabhai Prajapati, J. / Kleinekathofer, U. / Winterhalter, M. | ||||||
| History |
| ||||||
| Remark 700 | SHEET DETERMINATION METHOD: DSSP THE SHEETS PRESENTED AS "AA" IN EACH CHAIN ON SHEET RECORDS BELOW ... SHEET DETERMINATION METHOD: DSSP THE SHEETS PRESENTED AS "AA" IN EACH CHAIN ON SHEET RECORDS BELOW IS ACTUALLY AN 14-STRANDED BARREL THIS IS REPRESENTED BY A 15-STRANDED SHEET IN WHICH THE FIRST AND LAST STRANDS ARE IDENTICAL. SHEET DETERMINATION METHOD: DSSP THE SHEETS PRESENTED AS "BA" IN EACH CHAIN ON SHEET RECORDS BELOW IS ACTUALLY AN 14-STRANDED BARREL THIS IS REPRESENTED BY A 15-STRANDED SHEET IN WHICH THE FIRST AND LAST STRANDS ARE IDENTICAL. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4v3g.cif.gz 4v3g.cif.gz | 284.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4v3g.ent.gz pdb4v3g.ent.gz | 241.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4v3g.json.gz 4v3g.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  4v3g_validation.pdf.gz 4v3g_validation.pdf.gz | 2 MB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  4v3g_full_validation.pdf.gz 4v3g_full_validation.pdf.gz | 2 MB | Display | |
| Data in XML |  4v3g_validation.xml.gz 4v3g_validation.xml.gz | 26.5 KB | Display | |
| Data in CIF |  4v3g_validation.cif.gz 4v3g_validation.cif.gz | 35.3 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/v3/4v3g https://data.pdbj.org/pub/pdb/validation_reports/v3/4v3g ftp://data.pdbj.org/pub/pdb/validation_reports/v3/4v3g ftp://data.pdbj.org/pub/pdb/validation_reports/v3/4v3g | HTTPS FTP |
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 39999.426 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Details: CLONING REGION INCLUDING HEPTAHISTIDINE TAG AT N-TERMINUS Source: (gene. exp.)  KLEBSIELLA OXYTOCA (bacteria) / Plasmid: PB22 / Production host: KLEBSIELLA OXYTOCA (bacteria) / Plasmid: PB22 / Production host:  #2: Chemical | ChemComp-C8E / ( #3: Water | ChemComp-HOH / | Nonpolymer details | (HYDROXYETH | Sequence details | METHIONINE | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.2 Å3/Da / Density % sol: 62 % / Description: NONE |
|---|---|
| Crystal grow | pH: 8.5 Details: MOLECULAR DIMENSIONS MORPHEUS 1/35 OPTIMIZED, pH 8.5 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Diamond Diamond  / Beamline: I02 / Wavelength: 0.9796 / Beamline: I02 / Wavelength: 0.9796 |
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL / Date: Jun 22, 2013 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9796 Å / Relative weight: 1 |
| Reflection | Resolution: 2.5→50 Å / Num. obs: 32944 / % possible obs: 99.3 % / Observed criterion σ(I): -3 / Redundancy: 13.7 % / Biso Wilson estimate: 59.32 Å2 / Rmerge(I) obs: 0.08 / Net I/σ(I): 33.5 |
| Reflection shell | Resolution: 2.5→2.54 Å / Redundancy: 13.4 % / Rmerge(I) obs: 0.34 / Mean I/σ(I) obs: 5.7 / % possible all: 100 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  SAD SADStarting model: NONE Resolution: 2.513→48.998 Å / SU ML: 0.33 / σ(F): 0 / Phase error: 27.53 / Stereochemistry target values: ML
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.513→48.998 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller













 PDBj
PDBj




