[English] 日本語
 Yorodumi
Yorodumi- PDB-4kp5: Crystal structure of catalytic domain of human carbonic anhydrase... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4kp5 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of catalytic domain of human carbonic anhydrase isozyme XII with 2-Chloro-4-[(pyrimidin-2-ylsulfanyl)acetyl]benzenesulfonamide | ||||||
 Components Components | Carbonic anhydrase 12 | ||||||
 Keywords Keywords | LYASE/LYASE INHIBITOR / drug design / carbonic anhydrase / benzenesulfonamide / metal-binding / LYASE-LYASE INHIBITOR complex | ||||||
| Function / homology |  Function and homology information Function and homology informationchloride ion homeostasis / estrous cycle / Reversible hydration of carbon dioxide / carbonic anhydrase / carbonate dehydratase activity / basolateral plasma membrane / apical plasma membrane / zinc ion binding / membrane / plasma membrane Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 1.45 Å molecular replacement / Resolution: 1.45 Å | ||||||
 Authors Authors | Smirnov, A. / Manakova, E. / Grazulis, S. | ||||||
 Citation Citation |  Journal: Bioorg.Med.Chem. / Year: 2013 Journal: Bioorg.Med.Chem. / Year: 2013Title: Benzenesulfonamides with pyrimidine moiety as inhibitors of human carbonic anhydrases I, II, VI, VII, XII, and XIII Authors: Capkauskaite, E. / Zubriene, A. / Smirnov, A. / Torresan, J. / Kisonaite, M. / Kazokaite, J. / Gylyte, J. / Michailoviene, V. / Jogaite, V. / Manakova, E. / Grazulis, S. / Tumkevicius, S. / Matulis, D. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4kp5.cif.gz 4kp5.cif.gz | 255.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4kp5.ent.gz pdb4kp5.ent.gz | 205.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4kp5.json.gz 4kp5.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/kp/4kp5 https://data.pdbj.org/pub/pdb/validation_reports/kp/4kp5 ftp://data.pdbj.org/pub/pdb/validation_reports/kp/4kp5 ftp://data.pdbj.org/pub/pdb/validation_reports/kp/4kp5 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  4kniC  4knjC  4knmC  4knnC  4kp8C  1jd0S C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
-Protein , 1 types, 4 molecules ABCD
| #1: Protein | Mass: 29917.318 Da / Num. of mol.: 4 / Fragment: Catalytic domain, UNP residues 30-291 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: CA12 / Plasmid: pET21a / Production host: Homo sapiens (human) / Gene: CA12 / Plasmid: pET21a / Production host:  |
|---|
-Non-polymers , 5 types, 1221 molecules 
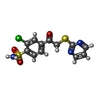







| #2: Chemical | ChemComp-ZN / #3: Chemical | ChemComp-E1F / #4: Chemical | ChemComp-EDO / #5: Chemical | ChemComp-SO4 / | #6: Water | ChemComp-HOH / | |
|---|
-Details
| Has protein modification | Y |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.1 Å3/Da / Density % sol: 40.7 % / Mosaicity: 0.23 ° |
|---|---|
| Crystal grow | Temperature: 291 K / Method: vapor diffusion, sitting drop / pH: 7 Details: 0.1M ammonium citrate pH 7.0, 0.2M ammonium sulfate, 30% PEG 4000, VAPOR DIFFUSION, SITTING DROP, temperature 291K |
-Data collection
| Diffraction | Mean temperature: 100 K | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  PETRA III, EMBL c/o DESY PETRA III, EMBL c/o DESY  / Beamline: P14 (MX2) / Wavelength: 0.826606 Å / Beamline: P14 (MX2) / Wavelength: 0.826606 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL / Date: Nov 30, 2012 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Monochromator: High Heat Load (HHL) Monochromator: Si 111; Large Offset Monochromator (LOM): Si 311, Si 511 Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 0.826606 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 0.945→86.515 Å / Num. obs: 168177 / % possible obs: 97.5 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 7 % / Biso Wilson estimate: 13.564 Å2 / Rmerge(I) obs: 0.045 / Rsym value: 0.045 / Net I/σ(I): 24.3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1
|
-Phasing
| Phasing | Method:  molecular replacement molecular replacement |
|---|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1JD0 Resolution: 1.45→73.13 Å / Cor.coef. Fo:Fc: 0.966 / Cor.coef. Fo:Fc free: 0.952 / WRfactor Rfree: 0.22 / WRfactor Rwork: 0.182 / Occupancy max: 1 / Occupancy min: 0.25 / FOM work R set: 0.848 / SU R Cruickshank DPI: 0.081 / SU Rfree: 0.084 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.081 / ESU R Free: 0.084 / Stereochemistry target values: MAXIMUM LIKELIHOOD Details: HYDROGENS HAVE BEEN USED IF PRESENT IN THE INPUT U VALUES : REFINED INDIVIDUALLY
| |||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: MASK | |||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 67.94 Å2 / Biso mean: 16.86 Å2 / Biso min: 2 Å2
| |||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.45→73.13 Å
| |||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.45→1.488 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller






















 PDBj
PDBj




