+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3seb | ||||||
|---|---|---|---|---|---|---|---|
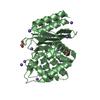
| Title | STAPHYLOCOCCAL ENTEROTOXIN B | ||||||
 Components Components | STAPHYLOCOCCAL ENTEROTOXIN B | ||||||
 Keywords Keywords | TOXIN / SUPERANTIGEN / ENTEROTOXIN | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.48 Å MOLECULAR REPLACEMENT / Resolution: 1.48 Å | ||||||
 Authors Authors | Papageorgiou, A.C. / Acharya, K.R. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1998 Journal: J.Mol.Biol. / Year: 1998Title: Crystal structure of microbial superantigen staphylococcal enterotoxin B at 1.5 A resolution: implications for superantigen recognition by MHC class II molecules and T-cell receptors. Authors: Papageorgiou, A.C. / Tranter, H.S. / Acharya, K.R. #1:  Journal: Structure / Year: 1997 Journal: Structure / Year: 1997Title: Superantigens as Immunomodulators: Recent Structural Insights Authors: Papageorgiou, A.C. / Acharya, K.R. #2:  Journal: Structure / Year: 1995 Journal: Structure / Year: 1995Title: Crystal Structure of the Superantigen Enterotoxin C2 from Staphylococcus Aureus Reveals a Zinc-Binding Site Authors: Papageorgiou, A.C. / Acharya, K.R. / Shapiro, R. / Passalacqua, E.F. / Brehm, R.D. / Tranter, H.S. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3seb.cif.gz 3seb.cif.gz | 64.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3seb.ent.gz pdb3seb.ent.gz | 47 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3seb.json.gz 3seb.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/se/3seb https://data.pdbj.org/pub/pdb/validation_reports/se/3seb ftp://data.pdbj.org/pub/pdb/validation_reports/se/3seb ftp://data.pdbj.org/pub/pdb/validation_reports/se/3seb | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 28281.885 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  |
|---|---|
| #2: Water | ChemComp-HOH / |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.15 Å3/Da / Density % sol: 42.75 % | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 8.5 / Details: pH 8.5 | ||||||||||||||||||||
| Crystal | *PLUS Density % sol: 2.05 % | ||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 16 ℃ / Method: vapor diffusion, hanging drop | ||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 289 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SRS SRS  / Beamline: PX9.6 / Wavelength: 0.87 / Beamline: PX9.6 / Wavelength: 0.87 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Jun 5, 1996 |
| Radiation | Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.87 Å / Relative weight: 1 |
| Reflection | Resolution: 1.5→20 Å / Num. obs: 38513 / % possible obs: 94.6 % / Observed criterion σ(I): 0 / Redundancy: 10 % / Rmerge(I) obs: 0.083 / Net I/σ(I): 12.5 |
| Reflection shell | Resolution: 1.5→1.55 Å / Redundancy: 3.5 % / Rmerge(I) obs: 0.364 / Mean I/σ(I) obs: 3.6 / % possible all: 79.9 |
| Reflection | *PLUS Num. measured all: 389081 |
| Reflection shell | *PLUS % possible obs: 79.9 % |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: SEB IN COMPLEX WITH AN MHC CLASS-II MOLECULE. Resolution: 1.48→20 Å / Num. parameters: 8551 / Num. restraintsaints: 7996 / Stereochemistry target values: ENGH AND HUBER Details: INITIAL REFINEMENT WAS CARRIED OUT WITH X-PLOR 3.1. THE LAST RECORDED R-FREE (5% OF THE REFLECTIONS EXCLUDED FROM THE REFINEMENT) WAS 0.252. AT THIS STAGE THE PROGRAM SHELXL-97 WAS EMPLOYED ...Details: INITIAL REFINEMENT WAS CARRIED OUT WITH X-PLOR 3.1. THE LAST RECORDED R-FREE (5% OF THE REFLECTIONS EXCLUDED FROM THE REFINEMENT) WAS 0.252. AT THIS STAGE THE PROGRAM SHELXL-97 WAS EMPLOYED FOR THE REMAINDER OF THE REFINEMENT. ALL DATA WERE USED. A POSTERIORI R-FREE CALCULATION GAVE AN R-FREE OF 0.209 WITH X-PLOR.
| |||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: MOEWS & KRETSINGER, J.MOL.BIOL.91(1973)201-228 | |||||||||||||||||||||||||||||||||
| Refine analyze | Num. disordered residues: 0 / Occupancy sum hydrogen: 0 / Occupancy sum non hydrogen: 2137 | |||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.48→20 Å
| |||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||
| Software | *PLUS Name: SHELXL-97 / Classification: refinement | |||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.182 | |||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller












 PDBj
PDBj

