+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3o7x | ||||||
|---|---|---|---|---|---|---|---|
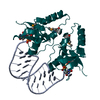
| Title | Crystal structure of human Hili PAZ domain | ||||||
 Components Components | Piwi-like protein 2 | ||||||
 Keywords Keywords | RNA BINDING PROTEIN / Piwi / RNA silencing / pi-RNA / Hiwi1 / Hili / PAZ domain | ||||||
| Function / homology |  Function and homology information Function and homology informationperinucleolar chromocenter / pi-body / PET complex / transposable element silencing by mRNA destabilization / secondary piRNA processing / positive regulation of meiosis I / piRNA binding / piRNA-mediated gene silencing by mRNA destabilization / transposable element silencing by piRNA-mediated heterochromatin formation / transposable element silencing by heterochromatin formation ...perinucleolar chromocenter / pi-body / PET complex / transposable element silencing by mRNA destabilization / secondary piRNA processing / positive regulation of meiosis I / piRNA binding / piRNA-mediated gene silencing by mRNA destabilization / transposable element silencing by piRNA-mediated heterochromatin formation / transposable element silencing by heterochromatin formation / piRNA processing / germ-line stem cell population maintenance / transposable element silencing by piRNA-mediated DNA methylation / negative regulation of circadian rhythm / chromatoid body / Hydrolases; Acting on ester bonds; Endoribonucleases producing 5'-phosphomonoesters / dense body / positive regulation of cytoplasmic translation / P granule / regulatory ncRNA-mediated gene silencing / oogenesis / PIWI-interacting RNA (piRNA) biogenesis / RNA endonuclease activity / meiotic cell cycle / rhythmic process / spermatogenesis / hydrolase activity / mRNA binding / metal ion binding / nucleus / cytoplasm Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  SAD / Resolution: 2.9248 Å SAD / Resolution: 2.9248 Å | ||||||
 Authors Authors | Tian, Y. / Simanshu, D.K. / Ma, J.-B. / Patel, D.J. | ||||||
 Citation Citation |  Journal: Proc.Natl.Acad.Sci.USA / Year: 2011 Journal: Proc.Natl.Acad.Sci.USA / Year: 2011Title: Inaugural Article: Structural basis for piRNA 2'-O-methylated 3'-end recognition by Piwi PAZ (Piwi/Argonaute/Zwille) domains. Authors: Tian, Y. / Simanshu, D.K. / Ma, J.B. / Patel, D.J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3o7x.cif.gz 3o7x.cif.gz | 101.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3o7x.ent.gz pdb3o7x.ent.gz | 78.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3o7x.json.gz 3o7x.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/o7/3o7x https://data.pdbj.org/pub/pdb/validation_reports/o7/3o7x ftp://data.pdbj.org/pub/pdb/validation_reports/o7/3o7x ftp://data.pdbj.org/pub/pdb/validation_reports/o7/3o7x | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | 
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 3 | 
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Unit cell |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Noncrystallographic symmetry (NCS) | NCS domain:
NCS domain segments:
|
- Components
Components
| #1: Protein | Mass: 16421.713 Da / Num. of mol.: 4 / Fragment: PAZ domain (UNP Residues 389-525) Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: PIWIL2, HILI / Plasmid: pET-28b / Production host: Homo sapiens (human) / Gene: PIWIL2, HILI / Plasmid: pET-28b / Production host:  #2: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.5 Å3/Da / Density % sol: 50.75 % |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 5.6 Details: 4-8% PEG4000 and 50 mM MgSO4 in 50 mM MES buffer, pH 5.6 to 6.1, VAPOR DIFFUSION, HANGING DROP, temperature 293K |
-Data collection
| Diffraction | Mean temperature: 100 K | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 24-ID-E / Wavelength: 0.9792 Å / Beamline: 24-ID-E / Wavelength: 0.9792 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Type: ADSC QUANTUM 315 / Detector: CCD Details: Cryogenically-cooled single crystal Si(111) side bounce monochromator | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 0.9792 Å / Relative weight: 1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Redundancy: 5.9 % / Av σ(I) over netI: 20.23 / Number: 78673 / Rmerge(I) obs: 0.081 / Χ2: 1.54 / D res high: 3 Å / D res low: 30 Å / Num. obs: 13333 / % possible obs: 97.5 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Diffraction reflection shell |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 2.92→50 Å / Num. obs: 14092 / % possible obs: 95.1 % / Redundancy: 6.9 % / Rmerge(I) obs: 0.091 / Χ2: 1.103 / Net I/σ(I): 20.2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell |
|
-Phasing
| Phasing | Method:  SAD SAD |
|---|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  SAD / Resolution: 2.9248→44.385 Å / Occupancy max: 1 / Occupancy min: 1 / SU ML: 0.37 / σ(F): 0.06 / Phase error: 32.41 / Stereochemistry target values: ML SAD / Resolution: 2.9248→44.385 Å / Occupancy max: 1 / Occupancy min: 1 / SU ML: 0.37 / σ(F): 0.06 / Phase error: 32.41 / Stereochemistry target values: ML
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL / Bsol: 41.941 Å2 / ksol: 0.32 e/Å3 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 135.33 Å2 / Biso mean: 75.2692 Å2 / Biso min: 29.25 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.9248→44.385 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints NCS |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Refine-ID: X-RAY DIFFRACTION / Total num. of bins used: 10
|
 Movie
Movie Controller
Controller













 PDBj
PDBj

