+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3hoe | ||||||
|---|---|---|---|---|---|---|---|
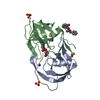
| Title | Crystal Structure of Surface Lipoprotein | ||||||
 Components Components | TbpB | ||||||
 Keywords Keywords | TRANSPORT PROTEIN / Iron acquisition / ion transport / receptor / transferrin / vaccine | ||||||
| Function / homology |  Function and homology information Function and homology informationLipocalin - #250 / Transferrin-binding protein B, C-lobe handle domain / TbpB, C-lobe handle domain superfamily / C-lobe handle domain of Tf-binding protein B / TbpB, N-lobe handle domain superfamily / Transferrin-binding protein B, C-lobe/N-lobe beta barrel domain / C-lobe and N-lobe beta barrels of Tf-binding protein B / Porin - #90 / Outer membrane protein/outer membrane enzyme PagP, beta-barrel / Porin ...Lipocalin - #250 / Transferrin-binding protein B, C-lobe handle domain / TbpB, C-lobe handle domain superfamily / C-lobe handle domain of Tf-binding protein B / TbpB, N-lobe handle domain superfamily / Transferrin-binding protein B, C-lobe/N-lobe beta barrel domain / C-lobe and N-lobe beta barrels of Tf-binding protein B / Porin - #90 / Outer membrane protein/outer membrane enzyme PagP, beta-barrel / Porin / Lipocalin / Prokaryotic membrane lipoprotein lipid attachment site profile. / Beta Barrel / Mainly Beta Similarity search - Domain/homology | ||||||
| Biological species |  Actinobacillus pleuropneumoniae (bacteria) Actinobacillus pleuropneumoniae (bacteria) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MAD / Resolution: 2.303 Å MAD / Resolution: 2.303 Å | ||||||
 Authors Authors | Moraes, T.F. / Yu, R.H. / Schryvers, A.B. / Strynadka, N.C.J. | ||||||
 Citation Citation |  Journal: Mol.Cell / Year: 2009 Journal: Mol.Cell / Year: 2009Title: Insights into the bacterial transferrin receptor: the structure of transferrin-binding protein B from Actinobacillus pleuropneumoniae. Authors: Moraes, T.F. / Yu, R.H. / Strynadka, N.C. / Schryvers, A.B. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3hoe.cif.gz 3hoe.cif.gz | 106.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3hoe.ent.gz pdb3hoe.ent.gz | 82.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3hoe.json.gz 3hoe.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ho/3hoe https://data.pdbj.org/pub/pdb/validation_reports/ho/3hoe ftp://data.pdbj.org/pub/pdb/validation_reports/ho/3hoe ftp://data.pdbj.org/pub/pdb/validation_reports/ho/3hoe | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 28101.988 Da / Num. of mol.: 2 / Fragment: residues 56-308 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Actinobacillus pleuropneumoniae (bacteria) Actinobacillus pleuropneumoniae (bacteria)Gene: tbpB / Production host:  #2: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.44 Å3/Da / Density % sol: 49.51 % |
|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, sitting drop / pH: 8 Details: 20% PEG3350, 0.1M Tris pH 8.0, 150 mM NaCl , VAPOR DIFFUSION, SITTING DROP, temperature 298K |
-Data collection
| Diffraction | Mean temperature: 100 K | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  CLSI CLSI  / Beamline: 08ID-1 / Wavelength: 0.98057, 0.98088, 0.97007 / Beamline: 08ID-1 / Wavelength: 0.98057, 0.98088, 0.97007 | ||||||||||||
| Detector | Type: MARMOSAIC 225 mm CCD / Detector: CCD / Date: Aug 13, 2008 Details: a double crystal monochromator (DCM) with an indirectly cryo-cooled first DCM w ith cryo-cooled 1st crystal sagittally bent 2nd crystal followed by vertically f ocusing mirror. | ||||||||||||
| Radiation | Monochromator: a double crystal monochromator (DCM) with an indirectly cryo-cooled first DCM with cryo-cooled 1st crystal sagittally bent 2nd crystal followed by vertically focusing mirror. Protocol: MAD / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||
| Radiation wavelength |
| ||||||||||||
| Reflection | Resolution: 2.303→35.31 Å / Num. obs: 42372 / % possible obs: 95 % / Observed criterion σ(F): 2 / Observed criterion σ(I): 2 / Redundancy: 5.9 % / Rsym value: 0.07 / Net I/σ(I): 29.6 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MAD / Resolution: 2.303→35.308 Å / Occupancy max: 1 / Occupancy min: 1 / SU ML: 0.33 / σ(F): 1.89 / Stereochemistry target values: ML MAD / Resolution: 2.303→35.308 Å / Occupancy max: 1 / Occupancy min: 1 / SU ML: 0.33 / σ(F): 1.89 / Stereochemistry target values: ML
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL / Bsol: 31.634 Å2 / ksol: 0.328 e/Å3 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 96.66 Å2 / Biso mean: 45.479 Å2 / Biso min: 22.82 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.303→35.308 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller













 PDBj
PDBj









