[English] 日本語
 Yorodumi
Yorodumi- PDB-3ad4: Crystal Structure of Methoxy Benzofuran Derivative bound to the K... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3ad4 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal Structure of Methoxy Benzofuran Derivative bound to the Kinase domain of human LCK, (auto-phosphorylated on TYR394) | ||||||
 Components Components | Proto-oncogene tyrosine-protein kinase LCK | ||||||
 Keywords Keywords | TRANSFERASE / TYROSINE-PROTEIN KINASE / ATP-BINDING / PHOSPHORYLATION / SIGNAL TRANSDUCTION / KINASE / SH2 DOMAIN / SH3 DOMAIN | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of lymphocyte activation / positive regulation of leukocyte cell-cell adhesion / CD27 signaling pathway / regulation of regulatory T cell differentiation / gamma-delta T cell differentiation / positive regulation of gamma-delta T cell differentiation / Fc-gamma receptor signaling pathway / FLT3 signaling through SRC family kinases / protein antigen binding / Nef Mediated CD4 Down-regulation ...regulation of lymphocyte activation / positive regulation of leukocyte cell-cell adhesion / CD27 signaling pathway / regulation of regulatory T cell differentiation / gamma-delta T cell differentiation / positive regulation of gamma-delta T cell differentiation / Fc-gamma receptor signaling pathway / FLT3 signaling through SRC family kinases / protein antigen binding / Nef Mediated CD4 Down-regulation / positive regulation of T cell receptor signaling pathway / CD4 receptor binding / Nef and signal transduction / Co-stimulation by CD28 / Interleukin-2 signaling / positive regulation of heterotypic cell-cell adhesion / CD28 dependent Vav1 pathway / leukocyte migration / Regulation of KIT signaling / phospholipase activator activity / Co-inhibition by CTLA4 / Translocation of ZAP-70 to Immunological synapse / Phosphorylation of CD3 and TCR zeta chains / PECAM1 interactions / protein serine/threonine phosphatase activity / pericentriolar material / CD8 receptor binding / hemopoiesis / Generation of second messenger molecules / T cell differentiation / immunological synapse / Co-inhibition by PD-1 / RHOH GTPase cycle / CD28 dependent PI3K/Akt signaling / phosphatidylinositol 3-kinase binding / GPVI-mediated activation cascade / phospholipase binding / T cell costimulation / T cell receptor binding / positive regulation of intrinsic apoptotic signaling pathway / phosphotyrosine residue binding / release of sequestered calcium ion into cytosol / SH2 domain binding / cell surface receptor protein tyrosine kinase signaling pathway / Signaling by phosphorylated juxtamembrane, extracellular and kinase domain KIT mutants / T cell activation / B cell receptor signaling pathway / non-specific protein-tyrosine kinase / non-membrane spanning protein tyrosine kinase activity / Signaling by SCF-KIT / platelet activation / positive regulation of T cell activation / Constitutive Signaling by Aberrant PI3K in Cancer / cell-cell junction / DAP12 signaling / PIP3 activates AKT signaling / Downstream TCR signaling / T cell receptor signaling pathway / ATPase binding / PI5P, PP2A and IER3 Regulate PI3K/AKT Signaling / protein tyrosine kinase activity / protein phosphatase binding / intracellular signal transduction / membrane raft / response to xenobiotic stimulus / signaling receptor binding / positive regulation of gene expression / protein kinase binding / extracellular exosome / ATP binding / identical protein binding / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.2 Å MOLECULAR REPLACEMENT / Resolution: 2.2 Å | ||||||
 Authors Authors | Tsuji, E. | ||||||
 Citation Citation |  Journal: To be Published Journal: To be PublishedTitle: Ab initio fragment molecular orbital study of ligand binding to leukocyte-specific protein tyrosine (LCK) kinase Authors: Ozawa, M. / Ozawa, T. / Tsuji, E. / Okazaki, K. / Takeda, K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3ad4.cif.gz 3ad4.cif.gz | 75.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3ad4.ent.gz pdb3ad4.ent.gz | 54.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3ad4.json.gz 3ad4.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ad/3ad4 https://data.pdbj.org/pub/pdb/validation_reports/ad/3ad4 ftp://data.pdbj.org/pub/pdb/validation_reports/ad/3ad4 ftp://data.pdbj.org/pub/pdb/validation_reports/ad/3ad4 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  3ac1C  3ac2C  3ac3C  3ac4C  3ac8C  3acjC  3ackC  3ad5C  3ad6C  3lckS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
-Protein , 1 types, 1 molecules A
| #1: Protein | Mass: 33065.602 Da / Num. of mol.: 1 / Fragment: RESIDUES 225-509 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: LCK / Plasmid: PET-19B / Cell line (production host): SF9 / Production host: Homo sapiens (human) / Gene: LCK / Plasmid: PET-19B / Cell line (production host): SF9 / Production host:  References: UniProt: P06239, non-specific protein-tyrosine kinase |
|---|
-Non-polymers , 5 types, 196 molecules 


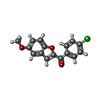





| #2: Chemical | | #3: Chemical | ChemComp-DMS / | #4: Chemical | ChemComp-MPD / ( | #5: Chemical | ChemComp-KBM / ( | #6: Water | ChemComp-HOH / | |
|---|
-Details
| Has protein modification | Y |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.18 Å3/Da / Density % sol: 43.65 % |
|---|---|
| Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop / pH: 6.5 Details: 0.2M (NH4)2SO4, 0.1M SODIUM CACODYLATE, 30% PEG 8000, 0.2% MPD, pH 6.5, VAPOR DIFFUSION, HANGING DROP, temperature 277K |
-Data collection
| Diffraction | Mean temperature: 93 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SPring-8 SPring-8  / Beamline: BL32B2 / Wavelength: 1.5418 Å / Beamline: BL32B2 / Wavelength: 1.5418 Å |
| Detector | Type: RIGAKU JUPITER 210 / Detector: CCD / Date: Nov 1, 2002 |
| Radiation | Monochromator: Si 111 CHANNEL / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.2→28.5 Å / Num. all: 14486 / Num. obs: 14511 / % possible obs: 95.4 % / Observed criterion σ(F): 0 / Redundancy: 3.7 % / Biso Wilson estimate: 15.9 Å2 / Rmerge(I) obs: 0.107 / Rsym value: 0.124 / Net I/σ(I): 12.5 |
| Reflection shell | Resolution: 2.2→2.32 Å / Redundancy: 3.9 % / Rmerge(I) obs: 0.213 / Mean I/σ(I) obs: 7.3 / Num. unique all: 2007 / Rsym value: 0.245 / % possible all: 92.2 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 3LCK Resolution: 2.2→10 Å / Cor.coef. Fo:Fc: 0.928 / Cor.coef. Fo:Fc free: 0.842 / SU B: 5.979 / SU ML: 0.155 / Cross valid method: THROUGHOUT / ESU R: 0.349 / ESU R Free: 0.259 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: BABINET MODEL WITH MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 20.559 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.2→10 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.2→2.254 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller




















 PDBj
PDBj



















