[English] 日本語
 Yorodumi
Yorodumi- PDB-1vbs: STRUCTURE OF CYCLOPHILIN COMPLEXED WITH (D)ALA CONTAINING TETRAPEPTIDE -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1vbs | ||||||
|---|---|---|---|---|---|---|---|
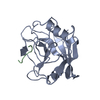
| Title | STRUCTURE OF CYCLOPHILIN COMPLEXED WITH (D)ALA CONTAINING TETRAPEPTIDE | ||||||
 Components Components |
| ||||||
 Keywords Keywords | ISOMERASE/ ISOMERASE SUBSTRATE / CYCLOPHILIN A / PEPTIDYL-PROLYL ISOMERASE / COMPETITIVE INHIBITOR / COMPLEX (ISOMERASE-PEPTIDE) / ISOMERASE- ISOMERASE SUBSTRATE COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology information: / negative regulation of protein K48-linked ubiquitination / regulation of apoptotic signaling pathway / cell adhesion molecule production / lipid droplet organization / negative regulation of viral life cycle / heparan sulfate binding / regulation of viral genome replication / virion binding / leukocyte chemotaxis ...: / negative regulation of protein K48-linked ubiquitination / regulation of apoptotic signaling pathway / cell adhesion molecule production / lipid droplet organization / negative regulation of viral life cycle / heparan sulfate binding / regulation of viral genome replication / virion binding / leukocyte chemotaxis / negative regulation of stress-activated MAPK cascade / activation of protein kinase B activity / endothelial cell activation / Basigin interactions / protein peptidyl-prolyl isomerization / cyclosporin A binding / Minus-strand DNA synthesis / Plus-strand DNA synthesis / Uncoating of the HIV Virion / Early Phase of HIV Life Cycle / Integration of provirus / APOBEC3G mediated resistance to HIV-1 infection / negative regulation of protein phosphorylation / intracellular membrane-bounded organelle / viral release from host cell / Calcineurin activates NFAT / Binding and entry of HIV virion / negative regulation of protein kinase activity / negative regulation of oxidative stress-induced intrinsic apoptotic signaling pathway / positive regulation of viral genome replication / neutrophil chemotaxis / Gene and protein expression by JAK-STAT signaling after Interleukin-12 stimulation / : / positive regulation of protein secretion / peptidylprolyl isomerase / peptidyl-prolyl cis-trans isomerase activity / Assembly Of The HIV Virion / Budding and maturation of HIV virion / platelet activation / positive regulation of protein phosphorylation / platelet aggregation / integrin binding / neuron differentiation / SARS-CoV-1 activates/modulates innate immune responses / : / Platelet degranulation / protein folding / cellular response to oxidative stress / secretory granule lumen / vesicle / ficolin-1-rich granule lumen / positive regulation of MAPK cascade / focal adhesion / apoptotic process / Neutrophil degranulation / protein-containing complex / : / RNA binding / extracellular exosome / extracellular region / membrane / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2 Å MOLECULAR REPLACEMENT / Resolution: 2 Å | ||||||
 Authors Authors | Zhao, Y. / Chen, Y. / Schutkowski, M. / Fischer, G. / Ke, H. | ||||||
 Citation Citation |  Journal: FEBS Lett. / Year: 1998 Journal: FEBS Lett. / Year: 1998Title: Mapping the stereospecificity of peptidyl prolyl cis/trans isomerases. Authors: Schiene, C. / Reimer, U. / Schutkowski, M. / Fischer, G. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1vbs.cif.gz 1vbs.cif.gz | 56.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1vbs.ent.gz pdb1vbs.ent.gz | 40.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1vbs.json.gz 1vbs.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/vb/1vbs https://data.pdbj.org/pub/pdb/validation_reports/vb/1vbs ftp://data.pdbj.org/pub/pdb/validation_reports/vb/1vbs ftp://data.pdbj.org/pub/pdb/validation_reports/vb/1vbs | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 18036.504 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Cell line: XA-90 F' / Gene: CYCLOPHILIN / Cell line (production host): XA-90 F' / Gene (production host): CYCLOPHILIN / Production host: Homo sapiens (human) / Cell line: XA-90 F' / Gene: CYCLOPHILIN / Cell line (production host): XA-90 F' / Gene (production host): CYCLOPHILIN / Production host:  |
|---|---|
| #2: Protein/peptide | Mass: 524.568 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Details: D-ALA CONTAINING TETRAPEPTIDE |
| #3: Water | ChemComp-HOH / |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.4 Å3/Da / Density % sol: 55 % |
|---|---|
| Crystal grow | pH: 8.2 / Details: pH 8.2 |
-Data collection
| Diffraction | Mean temperature: 174 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RUH2R / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU RUH2R / Wavelength: 1.5418 |
| Detector | Type: RIGAKU RAXIS II / Detector: IMAGE PLATE |
| Radiation | Monochromator: GRAPHITE(002) / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2→50 Å / Num. obs: 15416 / % possible obs: 88 % / Observed criterion σ(I): 1 / Redundancy: 3.3 % / Rmerge(I) obs: 0.088 / Rsym value: 0.088 / Net I/σ(I): 5.3 |
| Reflection shell | Resolution: 2→2.1 Å / % possible all: 76 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: UNLIGATED CYCLOPHILIN A Resolution: 2→10 Å / σ(F): 1
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 28.3 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→10 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints NCS | NCS model details: NOT APPLIED | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.1 / Classification: refinement X-PLOR / Version: 3.1 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS σ(F): 1 / % reflection Rfree: 5 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller











 PDBj
PDBj












