[English] 日本語
 Yorodumi
Yorodumi- PDB-1tgb: CRYSTAL STRUCTURE OF BOVINE TRYPSINOGEN AT 1.8 ANGSTROMS RESOLUTI... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1tgb | ||||||
|---|---|---|---|---|---|---|---|
| Title | CRYSTAL STRUCTURE OF BOVINE TRYPSINOGEN AT 1.8 ANGSTROMS RESOLUTION. II. CRYSTALLOGRAPHIC REFINEMENT, REFINED CRYSTAL STRUCTURE AND COMPARISON WITH BOVINE TRYPSIN | ||||||
 Components Components | TRYPSINOGEN | ||||||
 Keywords Keywords | HYDROLASE ZYMOGEN (SERINE PROTEINASE) | ||||||
| Function / homology |  Function and homology information Function and homology informationtrypsin / serpin family protein binding / serine protease inhibitor complex / digestion / endopeptidase activity / serine-type endopeptidase activity / proteolysis / : / metal ion binding Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 1.8 Å X-RAY DIFFRACTION / Resolution: 1.8 Å | ||||||
 Authors Authors | Bode, W. / Fehlhammer, H. / Huber, R. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1977 Journal: J.Mol.Biol. / Year: 1977Title: Crystal structure of bovine trypsinogen at 1-8 A resolution. II. Crystallographic refinement, refined crystal structure and comparison with bovine trypsin. Authors: Fehlhammer, H. / Bode, W. / Huber, R. #1:  Journal: J.Mol.Biol. / Year: 1976 Journal: J.Mol.Biol. / Year: 1976Title: Crystal Structure of Bovine Trypsinogen at 1.8 Angstroms Resolution. I. Data Collection, Application of Patterson Search Techniques and Preliminary Structural Interpretation Authors: Bode, W. / Fehlhammer, H. / Huber, R. #2:  Journal: Acta Crystallogr.,Sect.B / Year: 1980 Journal: Acta Crystallogr.,Sect.B / Year: 1980Title: Low-Temperature Protein Crystallography. Temperature Factor, Mosaic Spread, Extinction and Diffuse Scattering in Two Examples. Bovine Trypsinogen and Fc Fragment Authors: Singh, T.P. / Bode, W. / Huber, R. #3:  Journal: Acc.Chem.Res. / Year: 1978 Journal: Acc.Chem.Res. / Year: 1978Title: Structural Basis of the Activation and Action of Trypsin Authors: Huber, R. / Bode, W. #4:  Journal: FEBS Lett. / Year: 1978 Journal: FEBS Lett. / Year: 1978Title: Crystal Structure Analysis and Refinement of Two Variants of Trigonal Trypsinogen Authors: Bode, W. / Huber, R. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1tgb.cif.gz 1tgb.cif.gz | 55.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1tgb.ent.gz pdb1tgb.ent.gz | 39.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1tgb.json.gz 1tgb.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/tg/1tgb https://data.pdbj.org/pub/pdb/validation_reports/tg/1tgb ftp://data.pdbj.org/pub/pdb/validation_reports/tg/1tgb ftp://data.pdbj.org/pub/pdb/validation_reports/tg/1tgb | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 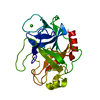
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Atom site foot note | 1: THESE ATOMS WERE NOT FOUND IN THE ELECTRON DENSITY MAP. THEIR COORDINATES WERE GENERATED USING STEREOCHEMICAL CRITERIA. | ||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 24012.953 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  |
|---|---|
| #2: Chemical | ChemComp-CA / |
| #3: Water | ChemComp-HOH / |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2 Å3/Da / Density % sol: 38.37 % |
|---|---|
| Crystal grow | *PLUS Method: other / Details: Bode, W., (1976) J. Mol. Biol., 106, 325. |
-Data collection
| Reflection | *PLUS Highest resolution: 1.8 Å / Num. obs: 15000 / Num. measured all: 45000 |
|---|
- Processing
Processing
| Refinement | Highest resolution: 1.8 Å Details: THE TEMPERATURE FACTOR FIELD OF THE ATOM AND HETATM RECORDS CONTAINS THE ATOMIC RADIUS WHICH HAS BEEN TENTATIVELY REFINED. THE OCCUPANCY FACTORS (WEIGHTS) HAVE BEEN REFINED ONLY FOR THE SOLVENT ATOMS. | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement step | Cycle: LAST / Highest resolution: 1.8 Å
| ||||||||||||
| Refinement | *PLUS σ(I): 2 / Rfactor all: 0.38 / Rfactor obs: 0.27 | ||||||||||||
| Solvent computation | *PLUS | ||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller





















 PDBj
PDBj




