+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1qjv | ||||||
|---|---|---|---|---|---|---|---|
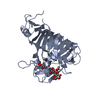
| Title | Pectin methylesterase PemA from Erwinia chrysanthemi | ||||||
 Components Components | PECTIN METHYLESTERASE | ||||||
 Keywords Keywords | HYDROLASE (ASPARTYL ESTERASE) / ESTERASE / PECTIN DEGRADATION / RIGHT-HANDED PARALLEL BETA HELIX | ||||||
| Function / homology |  Function and homology information Function and homology information: / pectinesterase / pectinesterase activity / cell wall modification / : / pectin catabolic process / cell outer membrane / : / extracellular region Similarity search - Function | ||||||
| Biological species |  ERWINIA CHRYSANTHEMI (bacteria) ERWINIA CHRYSANTHEMI (bacteria) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MIR / Resolution: 2.37 Å MIR / Resolution: 2.37 Å | ||||||
 Authors Authors | Jenkins, J. / Mayans, O. / Smith, D. / Worboys, K. / Pickersgill, R. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2001 Journal: J.Mol.Biol. / Year: 2001Title: Three-Dimensional Structure of Erwinia Chrysanthemi Pectin Methylesterase Reveals a Novel Esterase Active Site Authors: Jenkins, J. / Mayans, O. / Smith, D. / Worboys, K. / Pickersgill, R. #1: Journal: Gene / Year: 1993 Title: Characterization and Overexpression of the Pem Gene Encoding Pectin Methylesterase from Erwinia Chrysanthemi Strain-3937 Authors: Laurent, F. / Kotoujansky, A. / Labesse, G. / Bertheau, Y. #2: Journal: Appl.Microbiol.Biotechnol. / Year: 1991 Title: Production of Pectin Methylesterase from Erwinia Chrysanthemi B374 in Bacillus Subtilis Authors: Heikinheimo, R. / Hemila, H. / Pakkanen, R. / Palva, I. #3: Journal: Mol.Microbiol. / Year: 1988 Title: Molecular Cloning and Nucleotide Sequence of the Pectin Methyl Esterase Gene of Erwinia Chrysanthemi B374 Authors: Plastow, G.S. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1qjv.cif.gz 1qjv.cif.gz | 154.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1qjv.ent.gz pdb1qjv.ent.gz | 119.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1qjv.json.gz 1qjv.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qj/1qjv https://data.pdbj.org/pub/pdb/validation_reports/qj/1qjv ftp://data.pdbj.org/pub/pdb/validation_reports/qj/1qjv ftp://data.pdbj.org/pub/pdb/validation_reports/qj/1qjv | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (1, -0.00037, 0.00198), Vector: Details | BIOLOGICAL_UNIT: MONOMER | |
- Components
Components
| #1: Protein | Mass: 36963.773 Da / Num. of mol.: 2 / Fragment: MATURE ENZYME (RESIDUES 25-366) Source method: isolated from a genetically manipulated source Source: (gene. exp.)  ERWINIA CHRYSANTHEMI (bacteria) / Strain: B374 ERWINIA CHRYSANTHEMI (bacteria) / Strain: B374Description: THE EXPRESSION SYSTEM STRAIN IS FROM THE UK NATIONAL COLLECTION OF PLANT PATHOGENIC BACTERIA (NCPPB) Cellular location: EXTRACELLULAR / Gene: PEMA / Plasmid: PKTH1746 / Cellular location (production host): SECRETED / Gene (production host): ALPHA AMYLASE PROMOTER / Production host:  References: UniProt: P07863, UniProt: P0C1A8*PLUS, pectinesterase #2: Chemical | #3: Water | ChemComp-HOH / | Has protein modification | Y | Sequence details | STRAIN B374 BUT HISTIDINE 80 AS IN STRAIN 3937 COORDINATE | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.8 Å3/Da / Density % sol: 50 % | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Method: vapor diffusion, hanging drop / pH: 6.5 Details: HANGING DROP AGAINST 2.0 M AMMONIUM SULFATE AND 0.1M MES BUFFER AT PH 6.8 THE PROTEIN CONCENTATION WAS ABOUT 3 MG./ML. | |||||||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion, hanging drop | |||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  EMBL/DESY, HAMBURG EMBL/DESY, HAMBURG  / Beamline: X11 / Wavelength: 0.9058 / Beamline: X11 / Wavelength: 0.9058 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Oct 27, 1998 / Details: MIRRORS |
| Radiation | Monochromator: SI(111) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9058 Å / Relative weight: 1 |
| Reflection | Resolution: 2.4→50 Å / Num. obs: 32934 / % possible obs: 99 % / Redundancy: 4.04 % / Biso Wilson estimate: 14.6 Å2 / Rmerge(I) obs: 0.064 / Net I/σ(I): 13.43 |
| Reflection shell | Resolution: 2.4→2.44 Å / Redundancy: 3.04 % / Rmerge(I) obs: 0.217 / Mean I/σ(I) obs: 3 / % possible all: 82.3 |
| Reflection | *PLUS % possible obs: 99.1 % |
| Reflection shell | *PLUS % possible obs: 82.6 % / Rmerge(I) obs: 0.224 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MIR / Resolution: 2.37→20 Å / Rfactor Rfree error: 0.006 / Data cutoff high absF: 138672009.62 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: MAXIMUM LIKELIHOOD MIR / Resolution: 2.37→20 Å / Rfactor Rfree error: 0.006 / Data cutoff high absF: 138672009.62 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: MAXIMUM LIKELIHOODDetails: IN EACH PROTEIN CHAIN RESIDUE CYS 192 WAS FOUND TO HAVE TWO DIFFERENT CONFORMATIONS WITH EQUAL OCCUPANCIES THAT ARE ESSENTIALLY RELATED BY A 120 DEG. ROTATION ABOUT CHI1. THE CYS 192 IN EACH ...Details: IN EACH PROTEIN CHAIN RESIDUE CYS 192 WAS FOUND TO HAVE TWO DIFFERENT CONFORMATIONS WITH EQUAL OCCUPANCIES THAT ARE ESSENTIALLY RELATED BY A 120 DEG. ROTATION ABOUT CHI1. THE CYS 192 IN EACH CHAIN WITH ALTCODE B FORMS A DISULFIDE TO CYS 212. THE RESIDUE CYS 212 HAS ALMOST THE SAME CONFORMATION WITH AND WITHOUT THE DISULFIDE.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 51.2648 Å2 / ksol: 0.383205 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 17.9 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.37→20 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints NCS | Rms dev Biso : 0.701 Å2 / Rms dev position: 0.0243 Å / Weight Biso : 0.5 / Weight position: 400 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.37→2.52 Å / Rfactor Rfree error: 0.017 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: CNS / Version: 0.5 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor Rfree: 0.211 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS Biso mean: 17.8 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller













 PDBj
PDBj


