+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1oj6 | ||||||
|---|---|---|---|---|---|---|---|
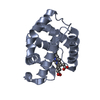
| Title | Human brain neuroglobin three-dimensional structure | ||||||
 Components Components | Neuroglobin | ||||||
 Keywords Keywords | OXYGEN TRANSPORT / NEUROGLOBIN / HEME HEXACOORDINATION | ||||||
| Function / homology |  Function and homology information Function and homology informationIntracellular oxygen transport / GDP-dissociation inhibitor activity / Oxidoreductases; Acting on other nitrogenous compounds as donors / nitrite reductase activity / oxygen transport / oxygen carrier activity / oxygen binding / cellular response to hypoxia / response to hypoxia / mitochondrial matrix ...Intracellular oxygen transport / GDP-dissociation inhibitor activity / Oxidoreductases; Acting on other nitrogenous compounds as donors / nitrite reductase activity / oxygen transport / oxygen carrier activity / oxygen binding / cellular response to hypoxia / response to hypoxia / mitochondrial matrix / heme binding / metal ion binding / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MAD / Resolution: 1.95 Å MAD / Resolution: 1.95 Å | ||||||
 Authors Authors | Pesce, A. / Dewilde, S. / Nardini, M. / Moens, L. / Ascenzi, P. / Hankeln, T. / Burmester, T. / Bolognesi, M. | ||||||
 Citation Citation |  Journal: Structure / Year: 2003 Journal: Structure / Year: 2003Title: Human Brain Neuroglobin Structure Reveals a Distinct Mode of Controlling Oxygen Affinity Authors: Pesce, A. / Dewilde, S. / Nardini, M. / Moens, L. / Ascenzi, P. / Hankeln, T. / Burmester, T. / Bolognesi, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1oj6.cif.gz 1oj6.cif.gz | 133.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1oj6.ent.gz pdb1oj6.ent.gz | 105.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1oj6.json.gz 1oj6.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/oj/1oj6 https://data.pdbj.org/pub/pdb/validation_reports/oj/1oj6 ftp://data.pdbj.org/pub/pdb/validation_reports/oj/1oj6 ftp://data.pdbj.org/pub/pdb/validation_reports/oj/1oj6 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| 4 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 16873.150 Da / Num. of mol.: 4 / Mutation: YES Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: NGB / Production host: Homo sapiens (human) / Gene: NGB / Production host:  #2: Chemical | ChemComp-HEM / #3: Chemical | ChemComp-SO4 / #4: Water | ChemComp-HOH / | Compound details | FUNCTION: INVOLVED IN OXYGEN TRANSPORT IN THE BRAIN. ENGINEERED MUTATION IN CHAINS A, B, C, D, CYS ...FUNCTION: INVOLVED IN OXYGEN TRANSPORT IN THE BRAIN. ENGINEERED | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.9 Å3/Da / Density % sol: 36 % | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 6.5 Details: 1.4 M AMMONIUM SULPHATE, 3% ISOPROPANOL, 0.05 M SODIUM CITRATE, PH 6.5 | ||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 277 K / pH: 6.5 / Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID29 / Wavelength: 1.738, 1.740, 0.933 / Beamline: ID29 / Wavelength: 1.738, 1.740, 0.933 | ||||||||||||
| Detector | Date: Mar 15, 2002 | ||||||||||||
| Radiation | Protocol: MAD / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||
| Radiation wavelength |
| ||||||||||||
| Reflection | Resolution: 1.95→40 Å / Num. obs: 35871 / % possible obs: 98.5 % / Redundancy: 2.6 % / Rmerge(I) obs: 0.055 / Net I/σ(I): 17 | ||||||||||||
| Reflection shell | Resolution: 1.95→1.98 Å / Redundancy: 2.5 % / Rmerge(I) obs: 0.22 / Mean I/σ(I) obs: 5 / % possible all: 97.9 | ||||||||||||
| Reflection | *PLUS Highest resolution: 1.95 Å / Lowest resolution: 40 Å / Redundancy: 2.6 % / Num. measured all: 92865 / Rmerge(I) obs: 0.055 | ||||||||||||
| Reflection shell | *PLUS % possible obs: 97.9 % / Redundancy: 2.5 % / Rmerge(I) obs: 0.22 / Mean I/σ(I) obs: 5 |
- Processing
Processing
| Software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MAD / Resolution: 1.95→40 Å / SU B: 4.111 / SU ML: 0.12 / Cross valid method: THROUGHOUT / ESU R: 0.216 / ESU R Free: 0.18 / Details: SOME ATOMS IN THIS ENTRY HAVE OCCUPANCY OF 0.00 MAD / Resolution: 1.95→40 Å / SU B: 4.111 / SU ML: 0.12 / Cross valid method: THROUGHOUT / ESU R: 0.216 / ESU R Free: 0.18 / Details: SOME ATOMS IN THIS ENTRY HAVE OCCUPANCY OF 0.00
| ||||||||||||||||||||
| Displacement parameters | Biso mean: 26 Å2 | ||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.95→40 Å
| ||||||||||||||||||||
| Refinement | *PLUS Lowest resolution: 40 Å / % reflection Rfree: 10 % | ||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller












 PDBj
PDBj












