[English] 日本語
 Yorodumi
Yorodumi- PDB-1o0p: Solution Structure of the third RNA Recognition Motif (RRM) of U2... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1o0p | ||||||
|---|---|---|---|---|---|---|---|
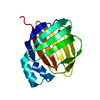
| Title | Solution Structure of the third RNA Recognition Motif (RRM) of U2AF65 in complex with an N-terminal SF1 peptide | ||||||
 Components Components |
| ||||||
 Keywords Keywords | RNA BINDING PROTEIN / NON-CANONICAL RNA RECOGNITION MOTIF / 4-STRANDED ANTI-PARALLEL BETA-SHEET / 2 ALPHA HELICES additionally extended by a third helix C | ||||||
| Function / homology |  Function and homology information Function and homology informationU2AF complex / poly-pyrimidine tract binding / pre-mRNA 3'-splice site binding / C2H2 zinc finger domain binding / mRNA 3'-end processing / commitment complex / mRNA cis splicing, via spliceosome / Transport of Mature mRNA derived from an Intron-Containing Transcript / RNA Polymerase II Transcription Termination / U2-type prespliceosome ...U2AF complex / poly-pyrimidine tract binding / pre-mRNA 3'-splice site binding / C2H2 zinc finger domain binding / mRNA 3'-end processing / commitment complex / mRNA cis splicing, via spliceosome / Transport of Mature mRNA derived from an Intron-Containing Transcript / RNA Polymerase II Transcription Termination / U2-type prespliceosome / molecular function inhibitor activity / mRNA 3'-splice site recognition / negative regulation of mRNA splicing, via spliceosome / spliceosomal complex assembly / Protein hydroxylation / mRNA Splicing - Major Pathway / negative regulation of protein ubiquitination / positive regulation of RNA splicing / spliceosomal complex / negative regulation of smooth muscle cell proliferation / mRNA splicing, via spliceosome / mRNA processing / transcription corepressor activity / nuclear speck / ribosome / mRNA binding / enzyme binding / RNA binding / zinc ion binding / nucleoplasm / identical protein binding / nucleus Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method | SOLUTION NMR / Restrained molecular dynamics using CNS, ARIA for ambiguous distance restraints | ||||||
 Authors Authors | Selenko, P. / Gregorovic, G. / Sprangers, R. / Stier, G. / Rhani, Z. / Kramer, A. / Sattler, M. | ||||||
 Citation Citation |  Journal: Mol.Cell / Year: 2003 Journal: Mol.Cell / Year: 2003Title: Structural Basis for the molecular recognition between human splicing factors U2AF65 and SF1/mBBP Authors: Selenko, P. / Gregorovic, G. / Sprangers, R. / Stier, G. / Rhani, Z. / Kramer, A. / Sattler, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1o0p.cif.gz 1o0p.cif.gz | 374 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1o0p.ent.gz pdb1o0p.ent.gz | 308.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1o0p.json.gz 1o0p.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/o0/1o0p https://data.pdbj.org/pub/pdb/validation_reports/o0/1o0p ftp://data.pdbj.org/pub/pdb/validation_reports/o0/1o0p ftp://data.pdbj.org/pub/pdb/validation_reports/o0/1o0p | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 11976.615 Da / Num. of mol.: 1 / Fragment: C-terminal RRM domain Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Plasmid: modified pET24d / Species (production host): Escherichia coli / Production host: Homo sapiens (human) / Plasmid: modified pET24d / Species (production host): Escherichia coli / Production host:  |
|---|---|
| #2: Protein/peptide | Mass: 1691.936 Da / Num. of mol.: 1 / Fragment: N-terminal peptide / Source method: obtained synthetically Details: THE PEPTIDE WAS CHEMICALLY SYNTHESIZED AND DERIVED FROM THE N-TERMINUS OF SF1. THE SEQUENCE OF THE PEPTIDE IS NATURALLY FOUND IN HOMO SAPIENS (HUMAN). References: GenBank: 2463198, UniProt: Q15637*PLUS |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Sample conditions | Ionic strength: 50mM salt / pH: 6.4 / Pressure: ambient / Temperature: 295 K | |||||||||
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Radiation wavelength | Relative weight: 1 | ||||||||||||||||||||
| NMR spectrometer |
|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: Restrained molecular dynamics using CNS, ARIA for ambiguous distance restraints Software ordinal: 1 Details: THE EXPERIMENTALLY DETERMINED DISTANCE (3232 NOEs, INCLUDING 258 FOR THE SF1 PEPTIDE, 64 INTERMOLECULAR, AND 2*35 FOR HYDROGEN BONDS) AND DIHEDRAL ANGLE RESTRAINTS (99) WERE APPLIED IN A ...Details: THE EXPERIMENTALLY DETERMINED DISTANCE (3232 NOEs, INCLUDING 258 FOR THE SF1 PEPTIDE, 64 INTERMOLECULAR, AND 2*35 FOR HYDROGEN BONDS) AND DIHEDRAL ANGLE RESTRAINTS (99) WERE APPLIED IN A MIXED TORSION AND CARTESIAN ANGLE DYNAMICS/SIMULATED ANNEALING PROTOCOL. STRUCTURAL QUALITY WAS ASSESED WITH PROCHECK. | ||||||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the least restraint violations,structures with the lowest energy Conformers calculated total number: 100 / Conformers submitted total number: 10 |
 Movie
Movie Controller
Controller












 PDBj
PDBj



 NMRPipe
NMRPipe