+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1lre | ||||||
|---|---|---|---|---|---|---|---|
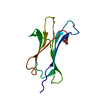
| Title | RECEPTOR ASSOCIATED PROTEIN (RAP) DOMAIN 1, NMR, 20 STRUCTURES | ||||||
 Components Components | RECEPTOR-ASSOCIATED PROTEIN | ||||||
 Keywords Keywords | CELL SURFACE PROTEIN / ALPHA2-MACROGLOBULIN RECEPTOR ASSOCIATED PROTEIN / LOW DENSITY LIPOPROTEIN RECEPTOR FAMILY ASSOCIATED PROTEIN / LDLR FAMILY ASSOCIATED PROTEIN / HELIX BUNDLE | ||||||
| Function / homology |  Function and homology information Function and homology informationextracellular negative regulation of signal transduction / regulation of receptor-mediated endocytosis / negative regulation of very-low-density lipoprotein particle clearance / rough endoplasmic reticulum lumen / lipase binding / receptor antagonist activity / amyloid-beta clearance by transcytosis / negative regulation of amyloid-beta clearance / very-low-density lipoprotein particle receptor binding / positive regulation of amyloid-beta clearance ...extracellular negative regulation of signal transduction / regulation of receptor-mediated endocytosis / negative regulation of very-low-density lipoprotein particle clearance / rough endoplasmic reticulum lumen / lipase binding / receptor antagonist activity / amyloid-beta clearance by transcytosis / negative regulation of amyloid-beta clearance / very-low-density lipoprotein particle receptor binding / positive regulation of amyloid-beta clearance / cis-Golgi network / negative regulation of receptor internalization / low-density lipoprotein particle receptor binding / endoplasmic reticulum-Golgi intermediate compartment / endomembrane system / endosome lumen / Golgi lumen / heparin binding / amyloid-beta binding / endosome / receptor ligand activity / signaling receptor binding / cell surface / endoplasmic reticulum / Golgi apparatus / signal transduction / extracellular region / plasma membrane Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method | SOLUTION NMR / DG, MD | ||||||
 Authors Authors | Nielsen, P.R. / Poulsen, F.M. | ||||||
 Citation Citation |  Journal: Proc.Natl.Acad.Sci.USA / Year: 1997 Journal: Proc.Natl.Acad.Sci.USA / Year: 1997Title: The solution structure of the N-terminal domain of alpha2-macroglobulin receptor-associated protein. Authors: Nielsen, P.R. / Ellgaard, L. / Etzerodt, M. / Thogersen, H.C. / Poulsen, F.M. #1:  Journal: To be Published Journal: To be PublishedTitle: Alpha2-Macroglobulin Receptor Associated Proteins Consist of Three Domains with Similar Structural Motifs. The 1H, 15N, and 13C Chemical Shift Assignments and Secondary Structure of the ...Title: Alpha2-Macroglobulin Receptor Associated Proteins Consist of Three Domains with Similar Structural Motifs. The 1H, 15N, and 13C Chemical Shift Assignments and Secondary Structure of the Truncated N-Terminal Domain of Human RAP Authors: Nielsen, P.R. / Ellgaard, L. / Jensen, P.H. / Thoegersen, H.C. / Poulsen, F.M. #2:  Journal: Eur.J.Biochem. / Year: 1997 Journal: Eur.J.Biochem. / Year: 1997Title: Dissection of the Domain Architecture of the Alpha2Macroglobulin-Receptor-Associated Protein Authors: Ellgaard, L. / Holtet, T.L. / Nielsen, P.R. / Etzerodt, M. / Gliemann, J. / Thogersen, H.C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1lre.cif.gz 1lre.cif.gz | 525 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1lre.ent.gz pdb1lre.ent.gz | 441.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1lre.json.gz 1lre.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/lr/1lre https://data.pdbj.org/pub/pdb/validation_reports/lr/1lre ftp://data.pdbj.org/pub/pdb/validation_reports/lr/1lre ftp://data.pdbj.org/pub/pdb/validation_reports/lr/1lre | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 9536.008 Da / Num. of mol.: 1 / Fragment: N-TERMINAL DOMAIN, DOMAIN 1, RESIDUES 17 - 97 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Cell line: BL21 / Plasmid: PT=7=H=6=UB / Production host: Homo sapiens (human) / Cell line: BL21 / Plasmid: PT=7=H=6=UB / Production host:  |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR |
|---|
- Sample preparation
Sample preparation
| Sample conditions | pH: 6.4 / Temperature: 298 K |
|---|---|
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer | Type: Bruker AMX / Manufacturer: Bruker / Model: AMX / Field strength: 600 MHz |
|---|
- Processing
Processing
| Software |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR software |
| ||||||||||||
| Refinement | Method: DG, MD / Software ordinal: 1 | ||||||||||||
| NMR ensemble | Conformer selection criteria: LOWEST ENERGY / Conformers calculated total number: 100 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller













 PDBj
PDBj

 X-PLOR
X-PLOR