+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1kve | ||||||
|---|---|---|---|---|---|---|---|
| Title | KILLER TOXIN FROM HALOTOLERANT YEAST | ||||||
 Components Components | (SMK TOXIN) x 2 | ||||||
 Keywords Keywords | TOXIN / HALOTOLERANT YEAST | ||||||
| Function / homology |  Function and homology information Function and homology informationtoxin sequestering activity / toxin activity / killing of cells of another organism / protein-containing complex / extracellular region Similarity search - Function | ||||||
| Biological species |  Pichia farinosa (fungus) Pichia farinosa (fungus) | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 1.8 Å X-RAY DIFFRACTION / Resolution: 1.8 Å | ||||||
 Authors Authors | Kashiwagi, T. / Kunishima, N. / Suzuki, C. / Tsuchiya, F. / Nikkuni, S. / Arata, Y. / Morikawa, K. | ||||||
 Citation Citation |  Journal: Structure / Year: 1997 Journal: Structure / Year: 1997Title: The novel acidophilic structure of the killer toxin from halotolerant yeast demonstrates remarkable folding similarity with a fungal killer toxin. Authors: Kashiwagi, T. / Kunishima, N. / Suzuki, C. / Tsuchiya, F. / Nikkuni, S. / Arata, Y. / Morikawa, K. #1:  Journal: To be Published Journal: To be PublishedTitle: Crystallization and Preliminary X-Ray Diffraction Studies of a Novel Killer Toxin from a Halotolerant Yeast Pichia Farinosa Authors: Kunishima, N. / Kashiwagi, T. / Suzuki, C. / Nikkuni, S. / Tsuchiya, F. / Arata, Y. / Morikawa, K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1kve.cif.gz 1kve.cif.gz | 66.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1kve.ent.gz pdb1kve.ent.gz | 50.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1kve.json.gz 1kve.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/kv/1kve https://data.pdbj.org/pub/pdb/validation_reports/kv/1kve ftp://data.pdbj.org/pub/pdb/validation_reports/kv/1kve ftp://data.pdbj.org/pub/pdb/validation_reports/kv/1kve | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||||||
| 2 | 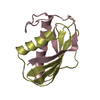
| ||||||||||||
| Unit cell |
| ||||||||||||
| Components on special symmetry positions |
| ||||||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (0.09656, -0.87326, 0.47759), Vector: |
- Components
Components
| #1: Protein | Mass: 6349.229 Da / Num. of mol.: 2 Fragment: CHAIN A AND C ARE RESIDUES 19 - 81 OF THE ALPHA CHAIN, CHAIN B AND D ARE RESIDUES 146 - 222 OF THE BETA CHAIN Source method: isolated from a natural source / Source: (natural)  Pichia farinosa (fungus) / Strain: KK1 / References: UniProt: P19972 Pichia farinosa (fungus) / Strain: KK1 / References: UniProt: P19972#2: Protein | Mass: 7856.556 Da / Num. of mol.: 2 Fragment: CHAIN A AND C ARE RESIDUES 19 - 81 OF THE ALPHA CHAIN, CHAIN B AND D ARE RESIDUES 146 - 222 OF THE BETA CHAIN Source method: isolated from a natural source / Source: (natural)  Pichia farinosa (fungus) / Strain: KK1 / References: UniProt: P19972 Pichia farinosa (fungus) / Strain: KK1 / References: UniProt: P19972#3: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.43 Å3/Da / Density % sol: 64.2 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 4 Details: CRYSTALS WERE GROWN FROM 25% POLYETHYLENE GLYCOL SOLUTION AT PH 4.0. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction source | Wavelength: 1.5418 |
|---|---|
| Detector | Type: MACSCIENCE / Detector: IMAGE PLATE / Date: Nov 20, 1995 |
| Radiation | Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 1.8→50 Å / Num. obs: 35468 / % possible obs: 94.7 % / Observed criterion σ(I): 0.5 / Redundancy: 5.23 % / Rmerge(I) obs: 0.062 |
| Reflection | *PLUS Num. measured all: 185448 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 1.8→6 Å / σ(F): 1 Details: IDEAL BOND LENGTHS AND ANGLES USED DURING REFINEMENT: HENDRICKSON AND KONNERT FINAL RMS COORD. SHIFT 0.007 ANGSTROMS INITIAL REFINEMENTS WERE DONE WITH X-PLOR 3.1 BY BRUNGER.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 22.49 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.15 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.8→6 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: PROLSQ / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.172 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller












 PDBj
PDBj
